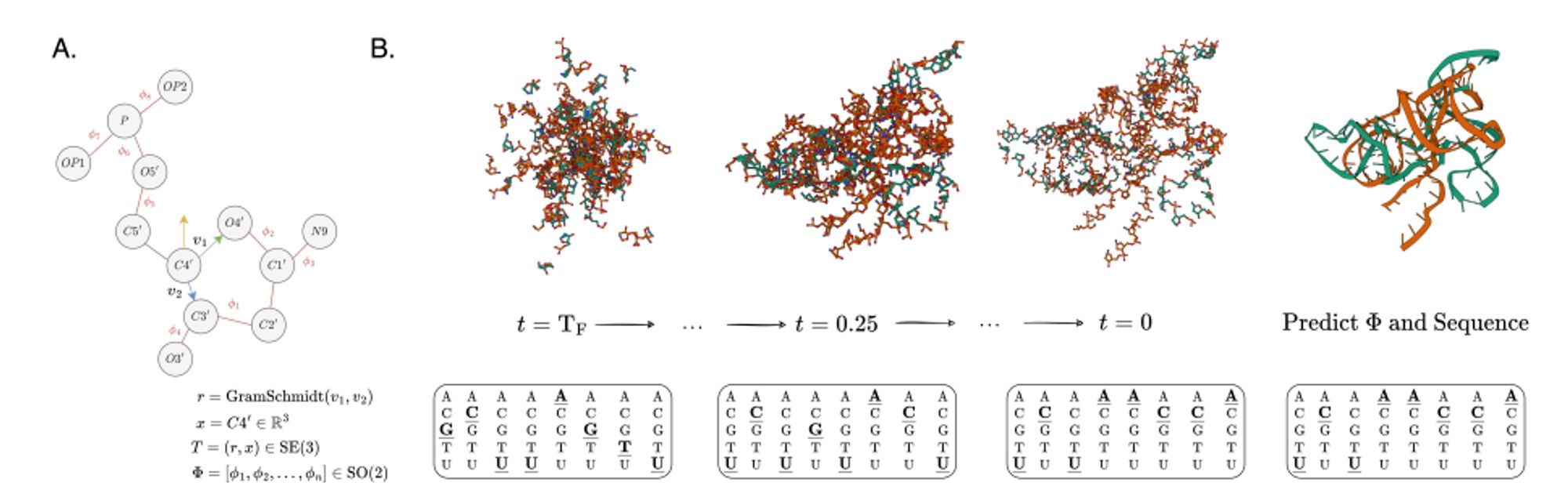
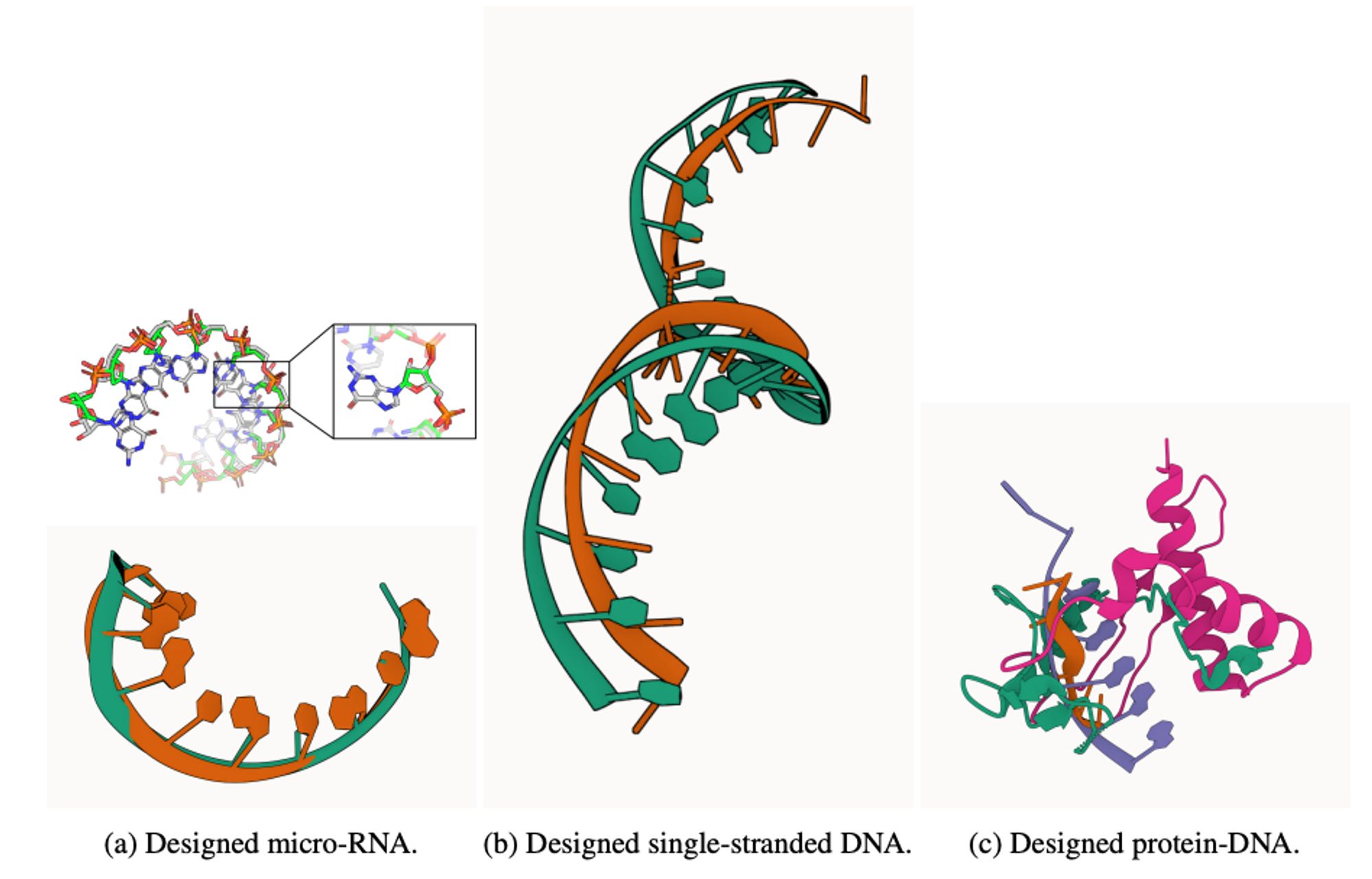
A diffusion model that simultaneously generates sequence and structure for protein-nucleic acid complexes. arxiv.org/abs/2401.06151
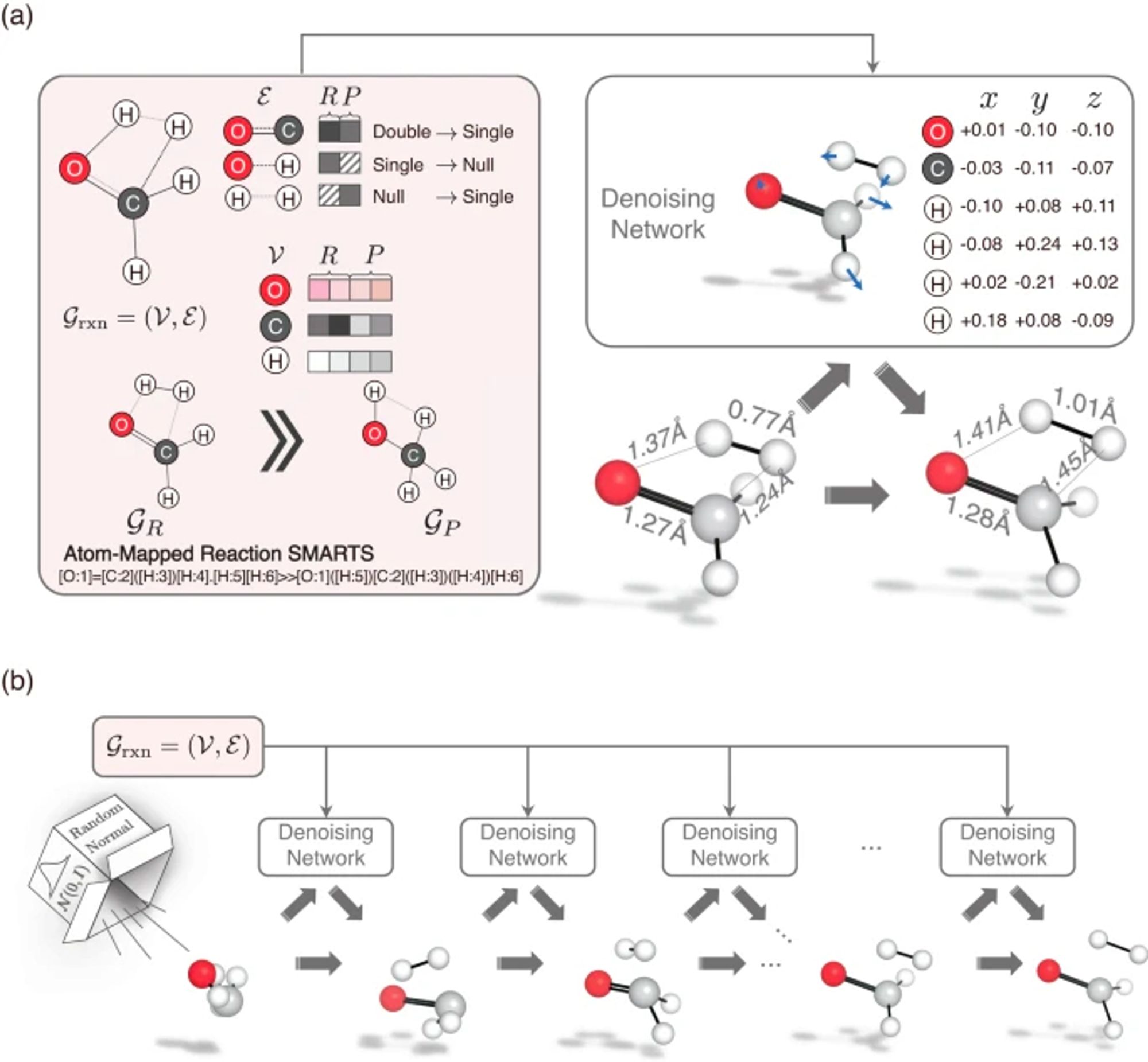
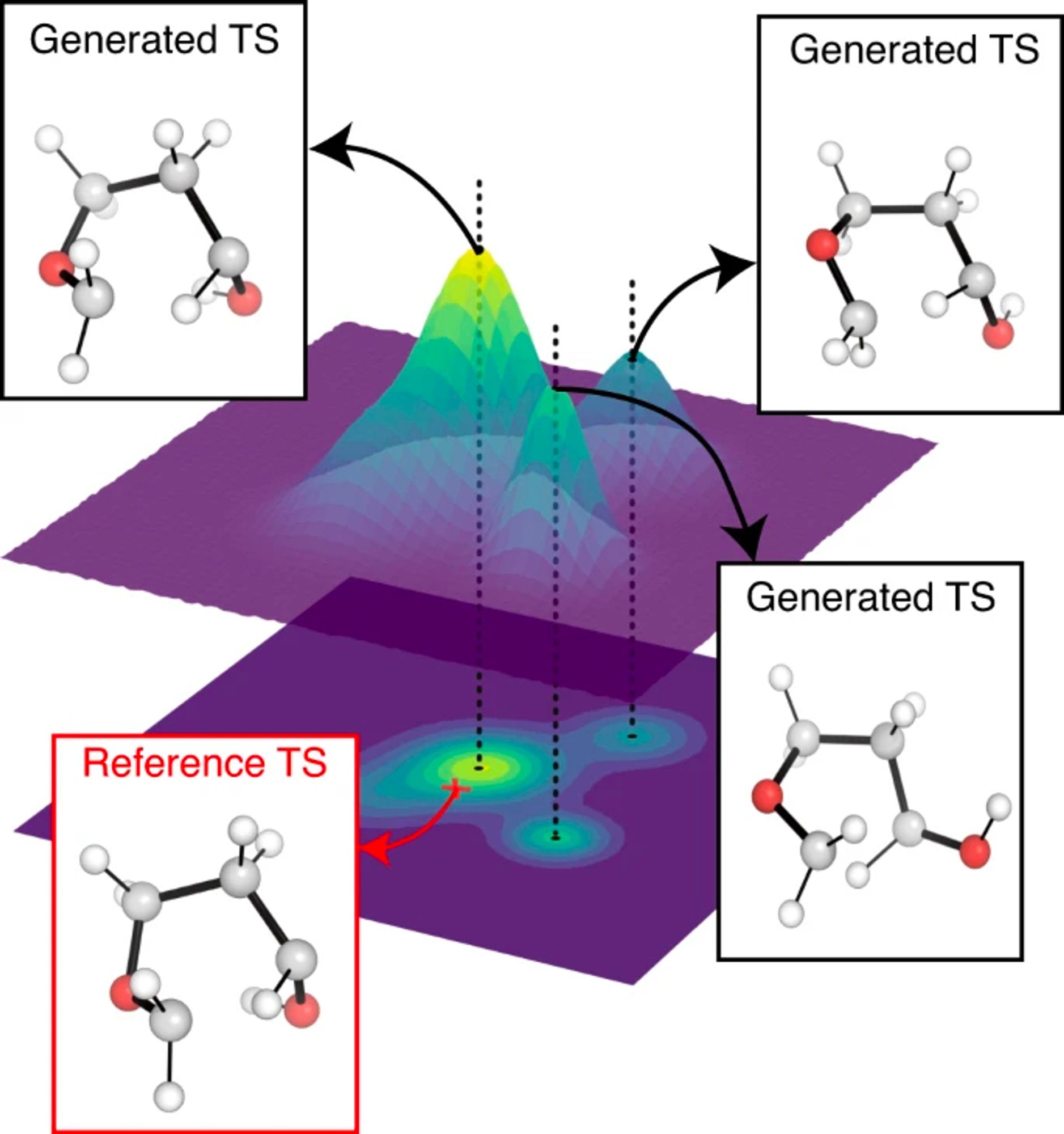
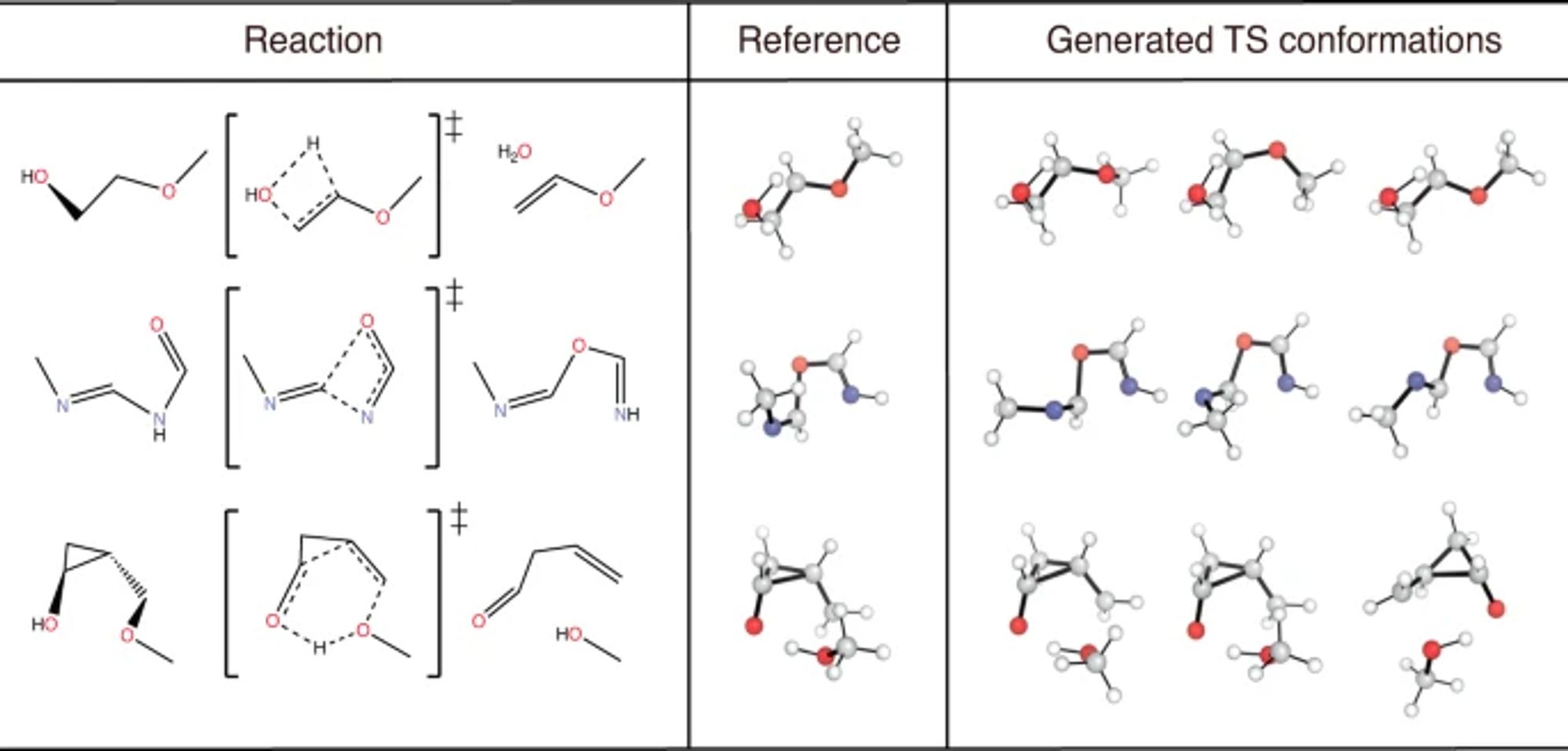
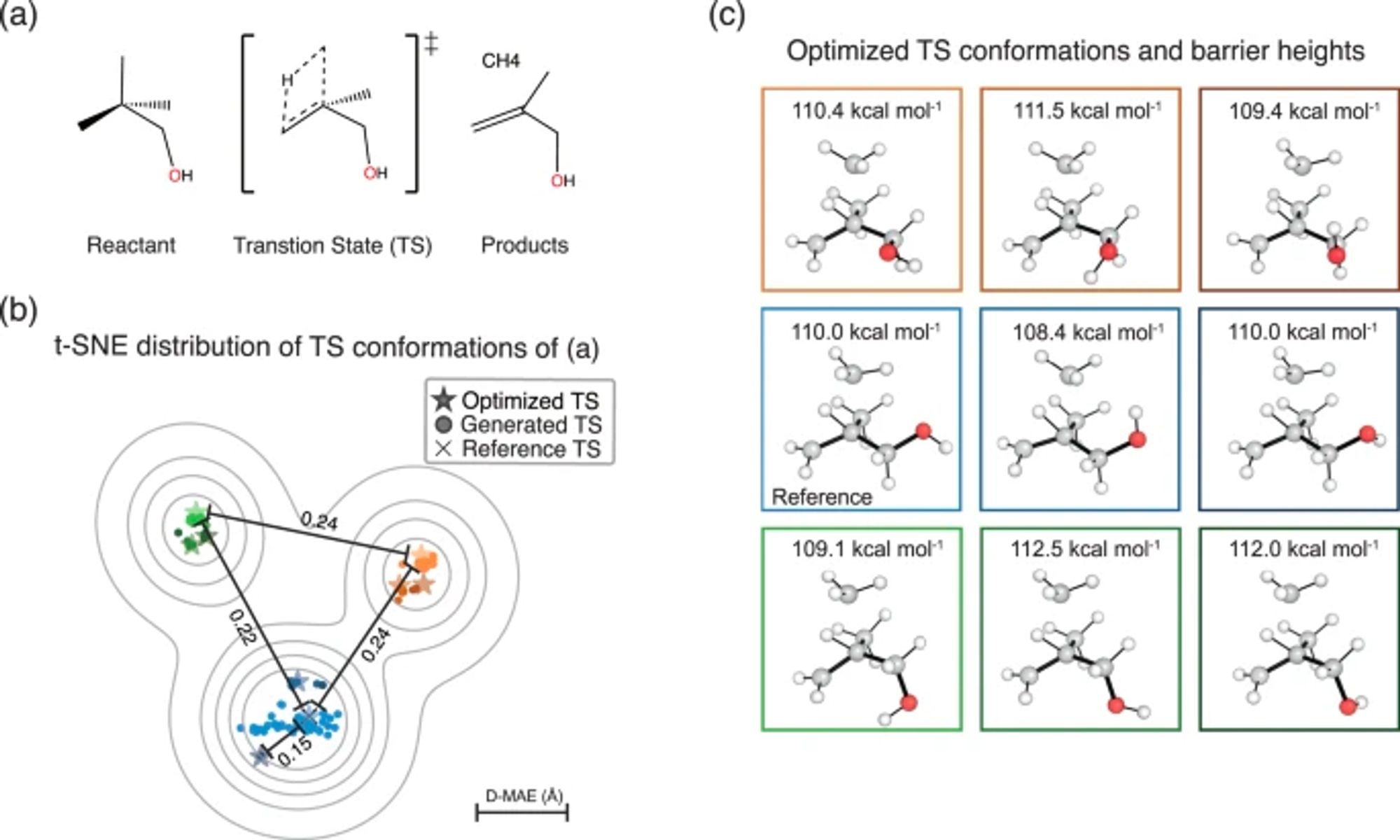
A graph diffusion model that predicts transition state geometries for chemical reactions www.nature.com/articles/s41...
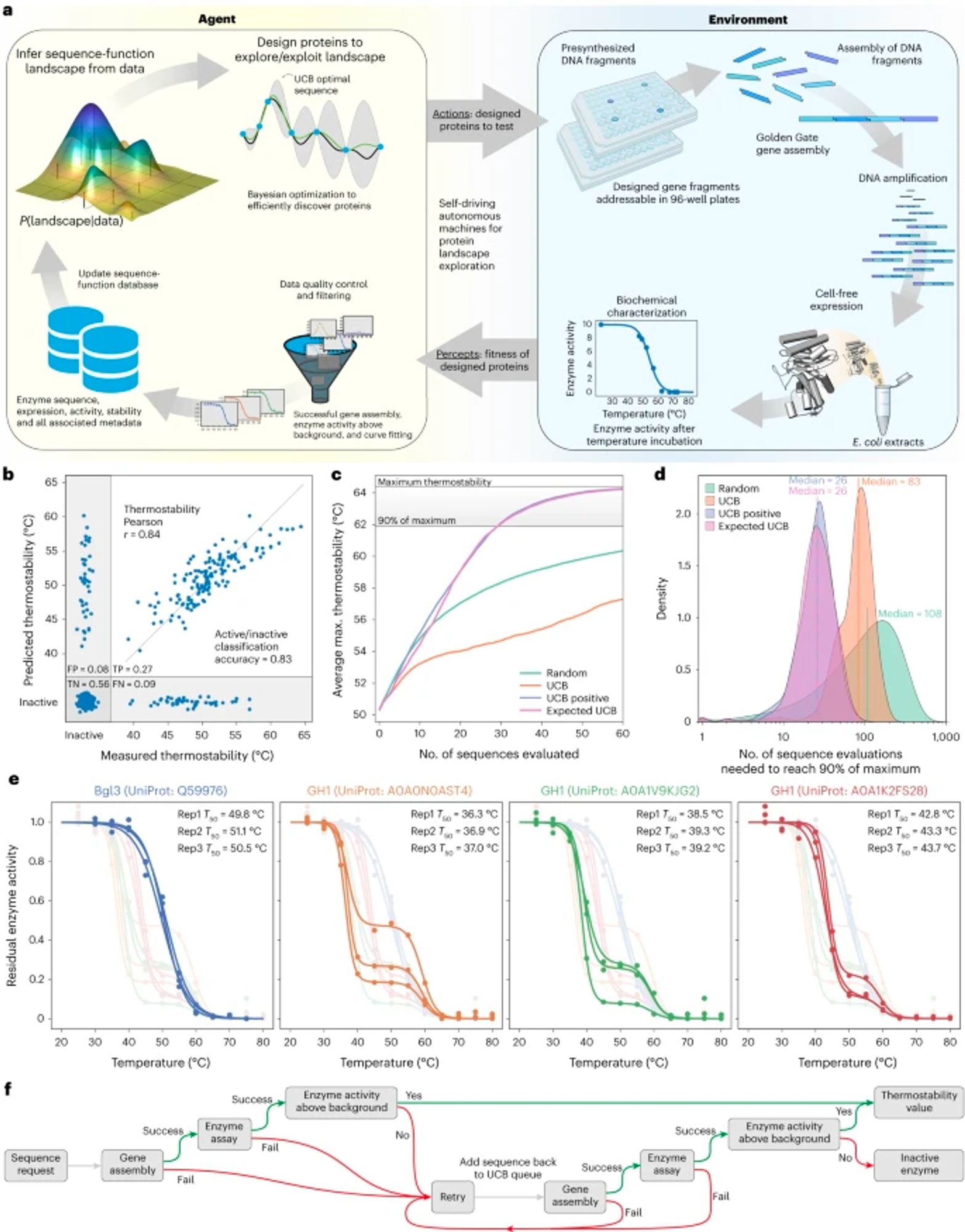
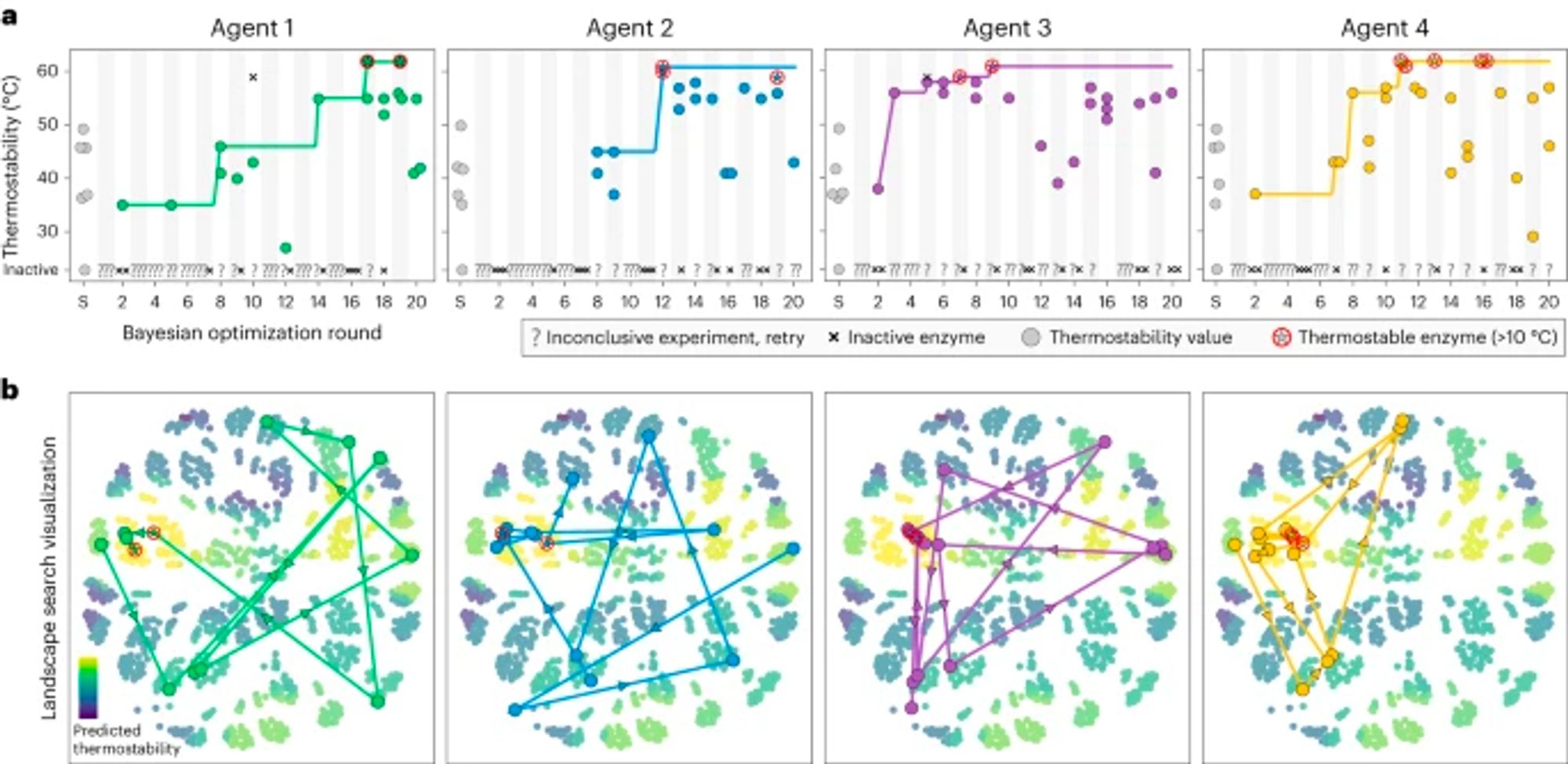
A robotic platform integrated with Bayesian optimization for autonomous protein engineering. "The procedure takes...9 h overall to go from a requested protein design to a physical protein sample to a corresponding data point." @romerolab1 www.nature.com/articles/s44...
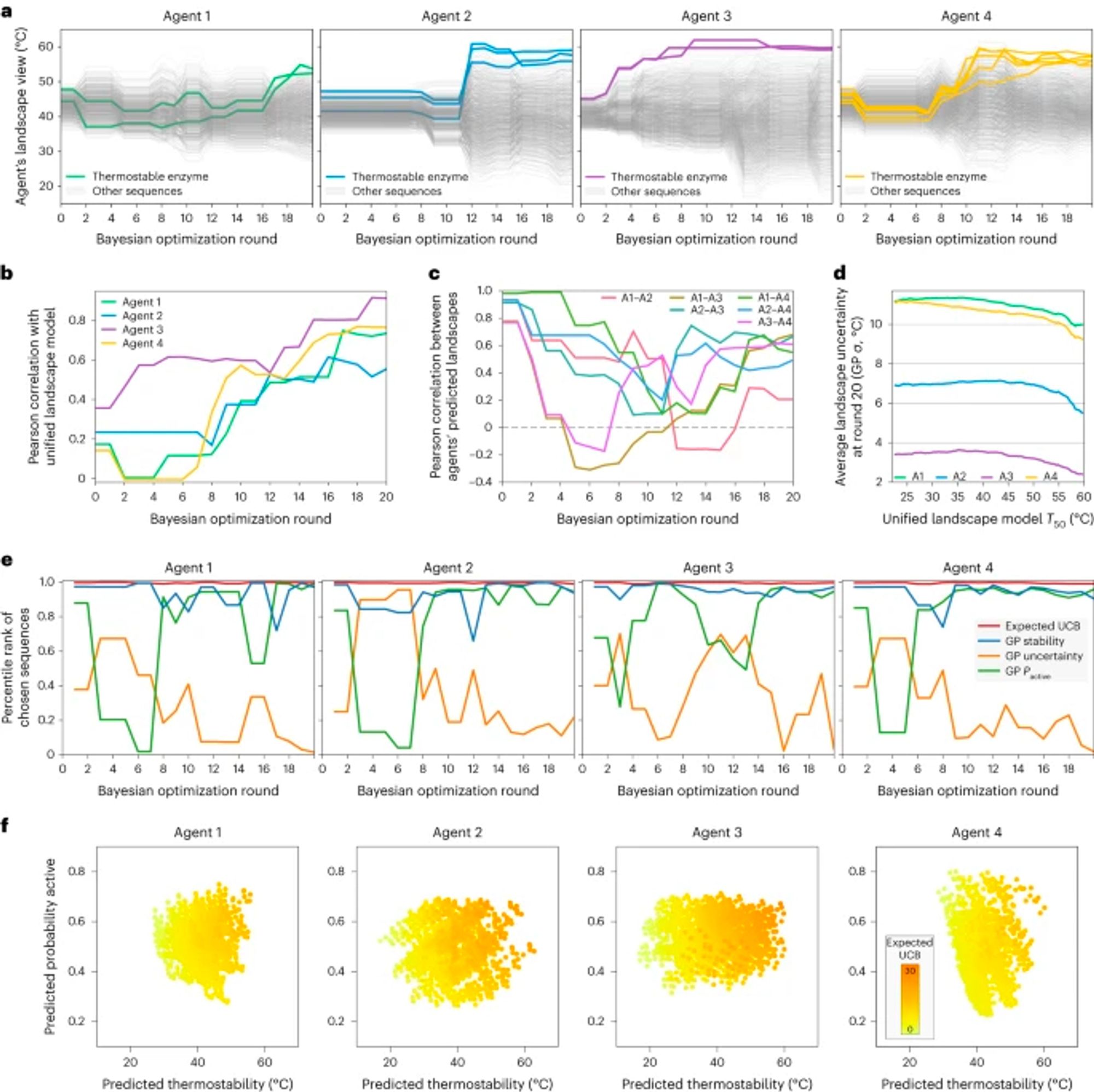
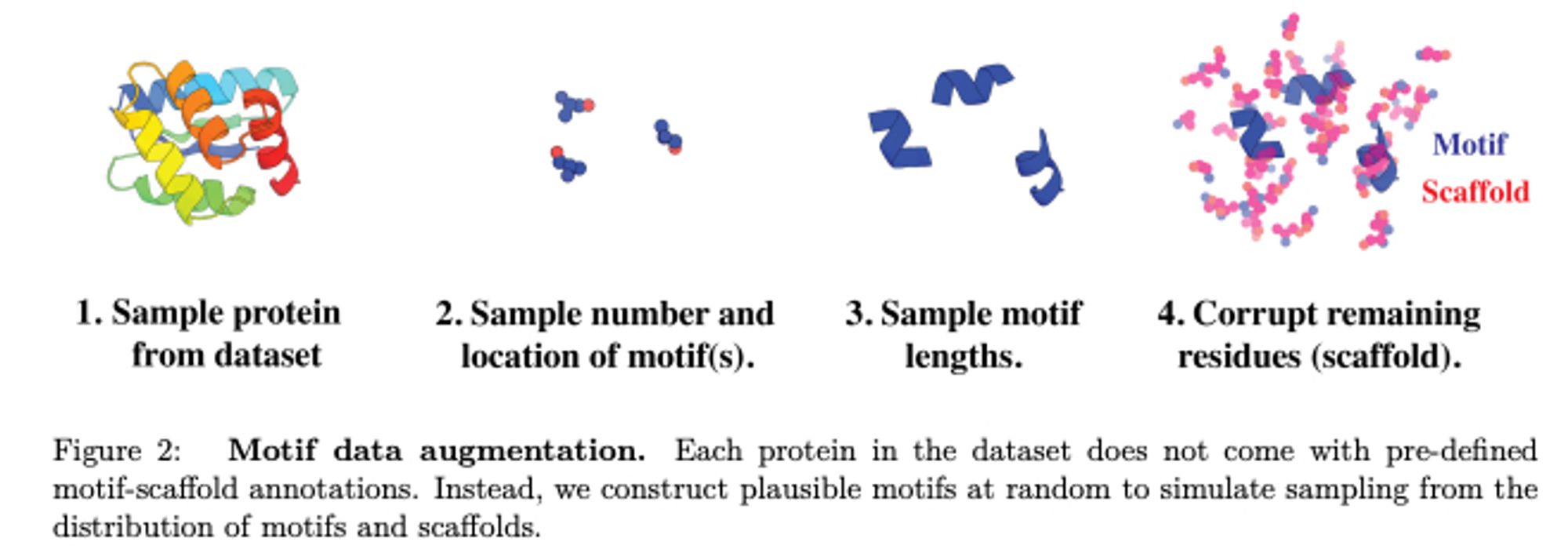
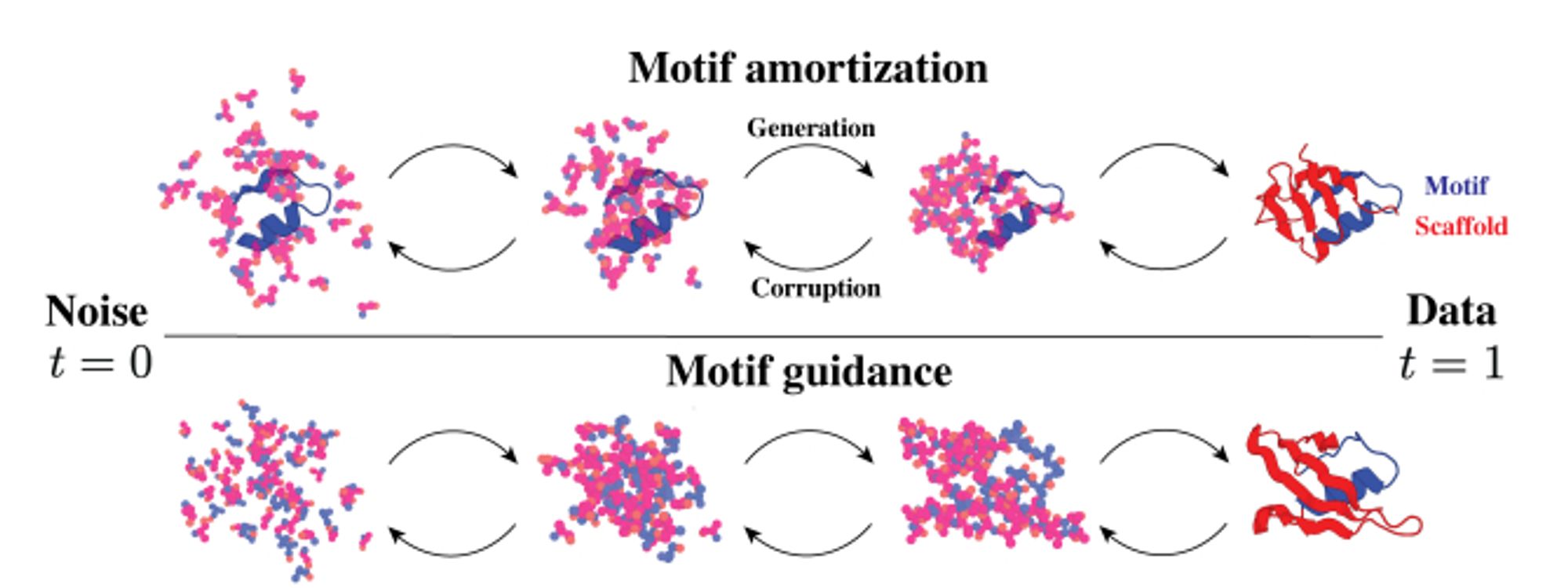
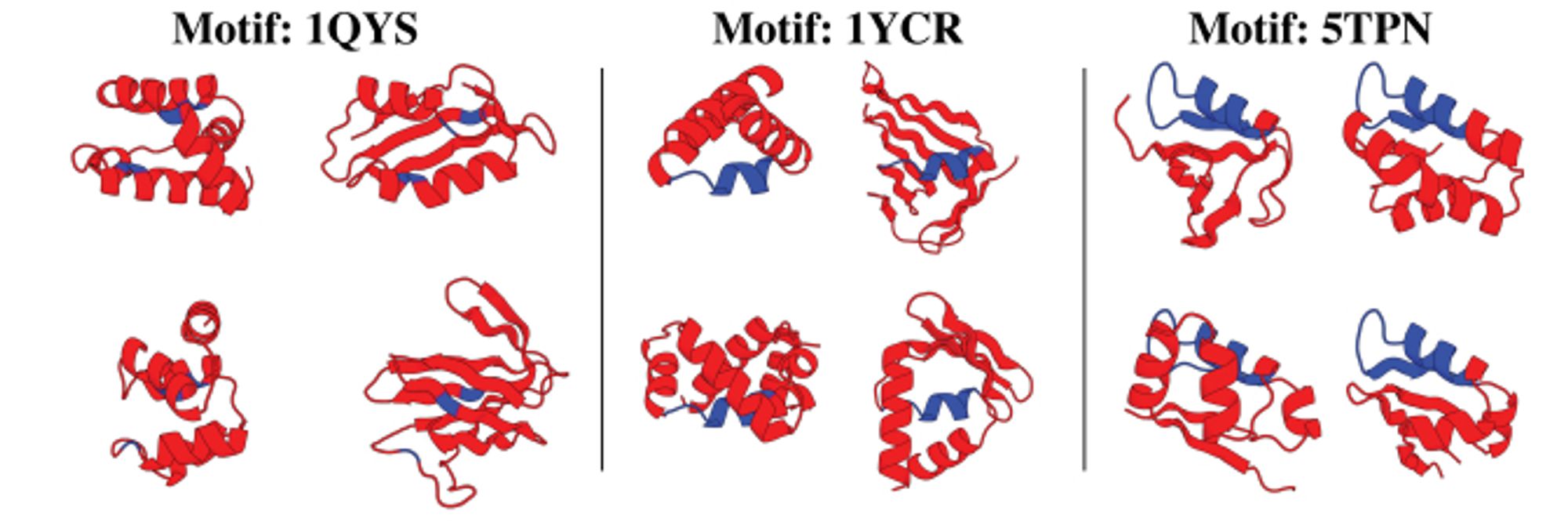
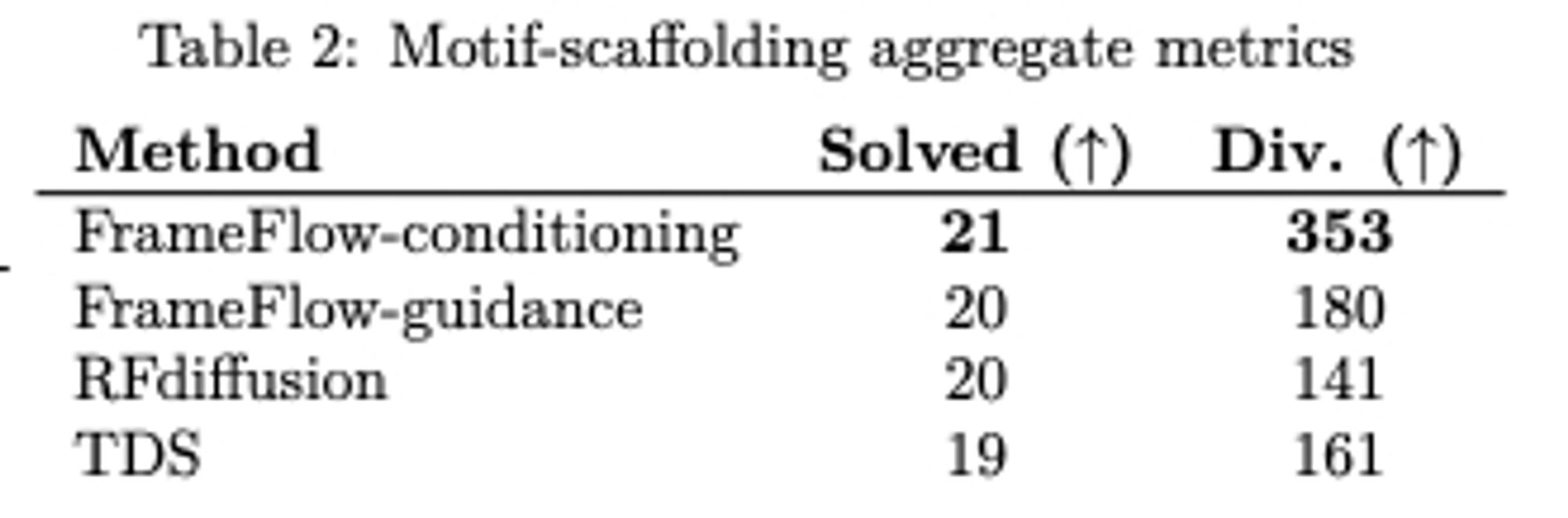
FrameDiff now extended to do motif scaffolding, with success rates close to RFdiffusion and better diversity. arxiv.org/abs/2401.04082

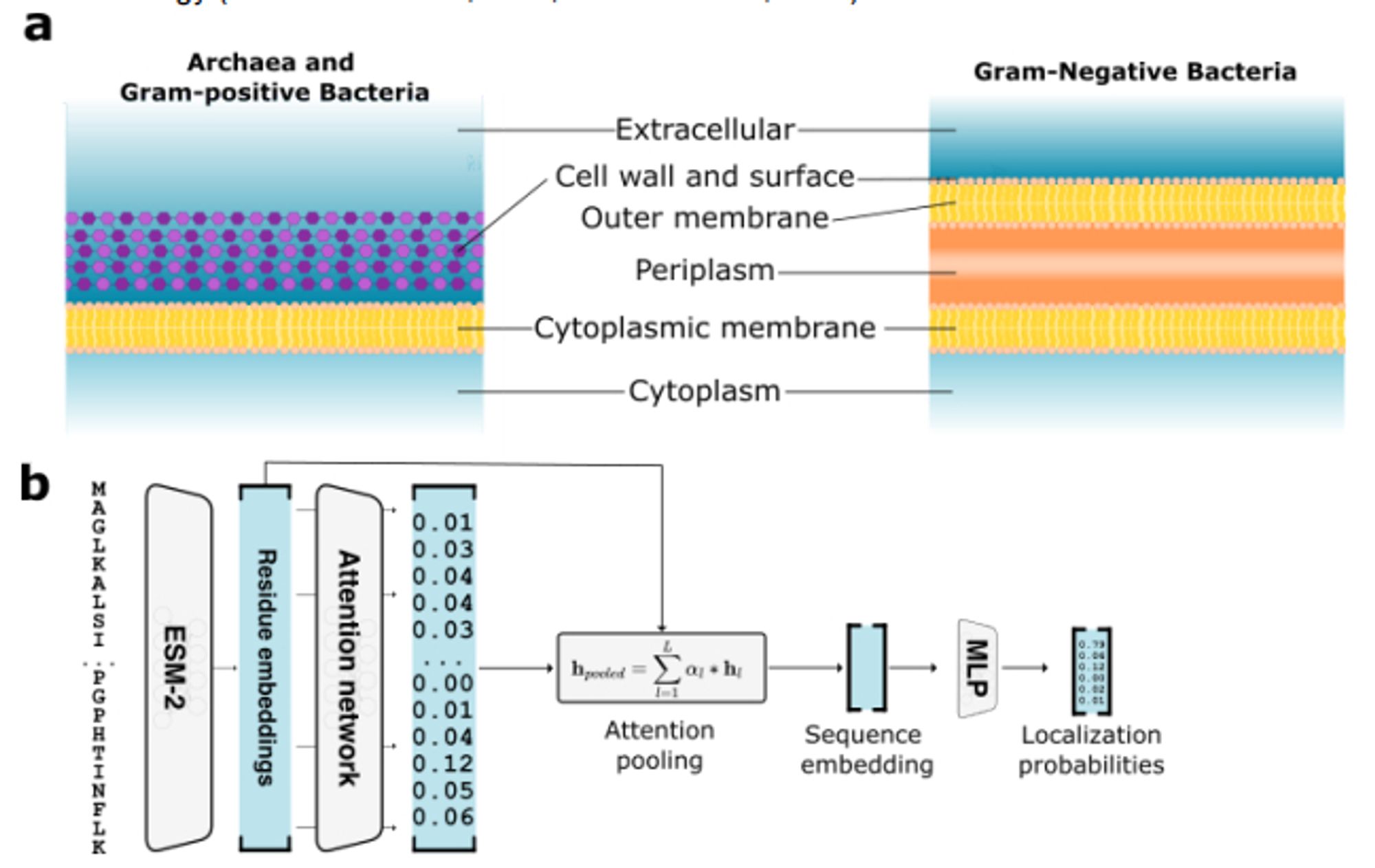
DeepLocPro: Use ESM-2 embeddings to predict prokaryotic protein subcellular localization. I'm kinda surprised that this is the first deep learning method for prokaryotic localization and that the SotA is from 2010! www.biorxiv.org/content/10.1...
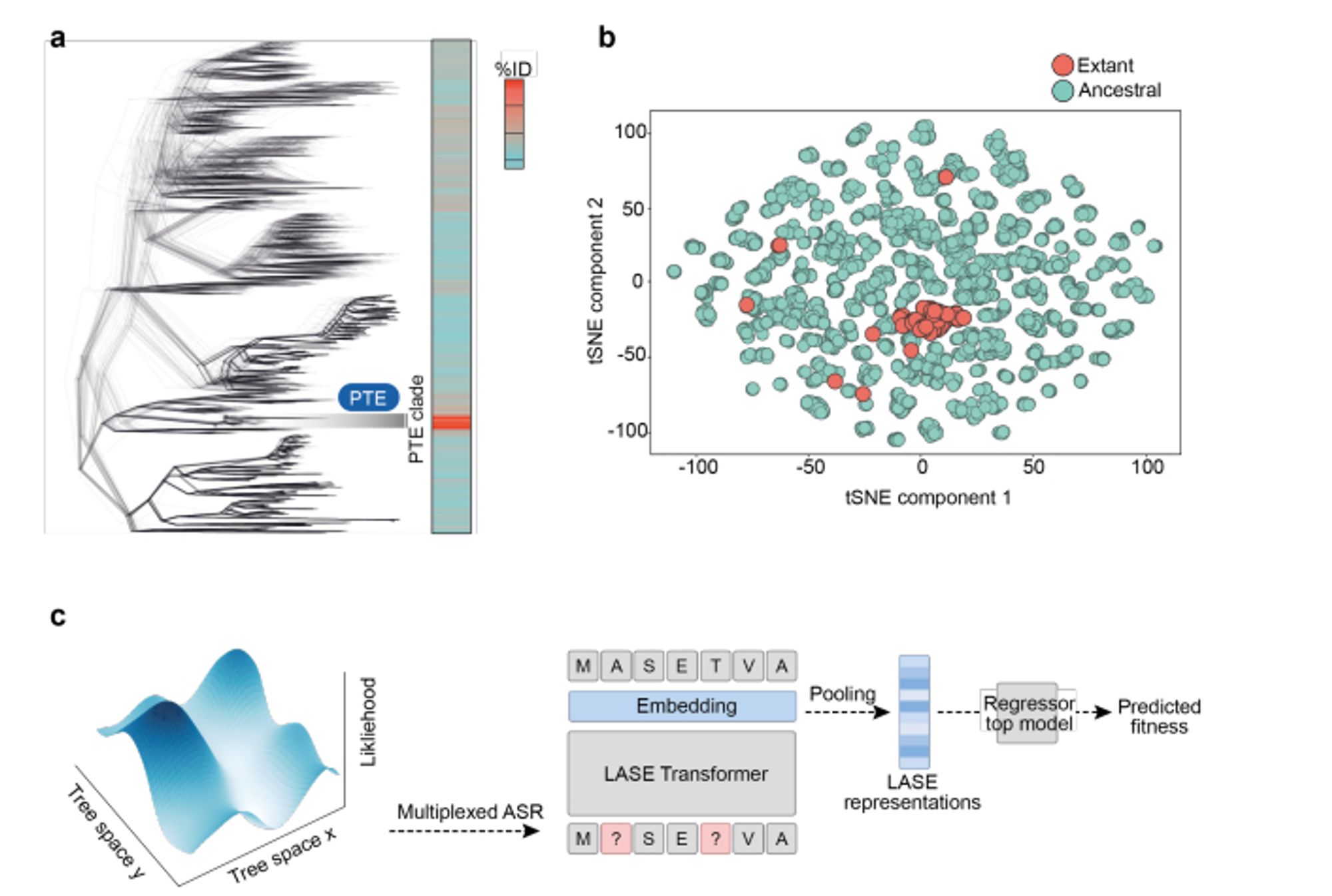
I spent a lot of time trying this when I first started at MSR, but I could never get it to work. The keys: - Generate many possible trees instead of just using the ancestral nodes from one. - Consider indels. www.biorxiv.org/content/10.1...
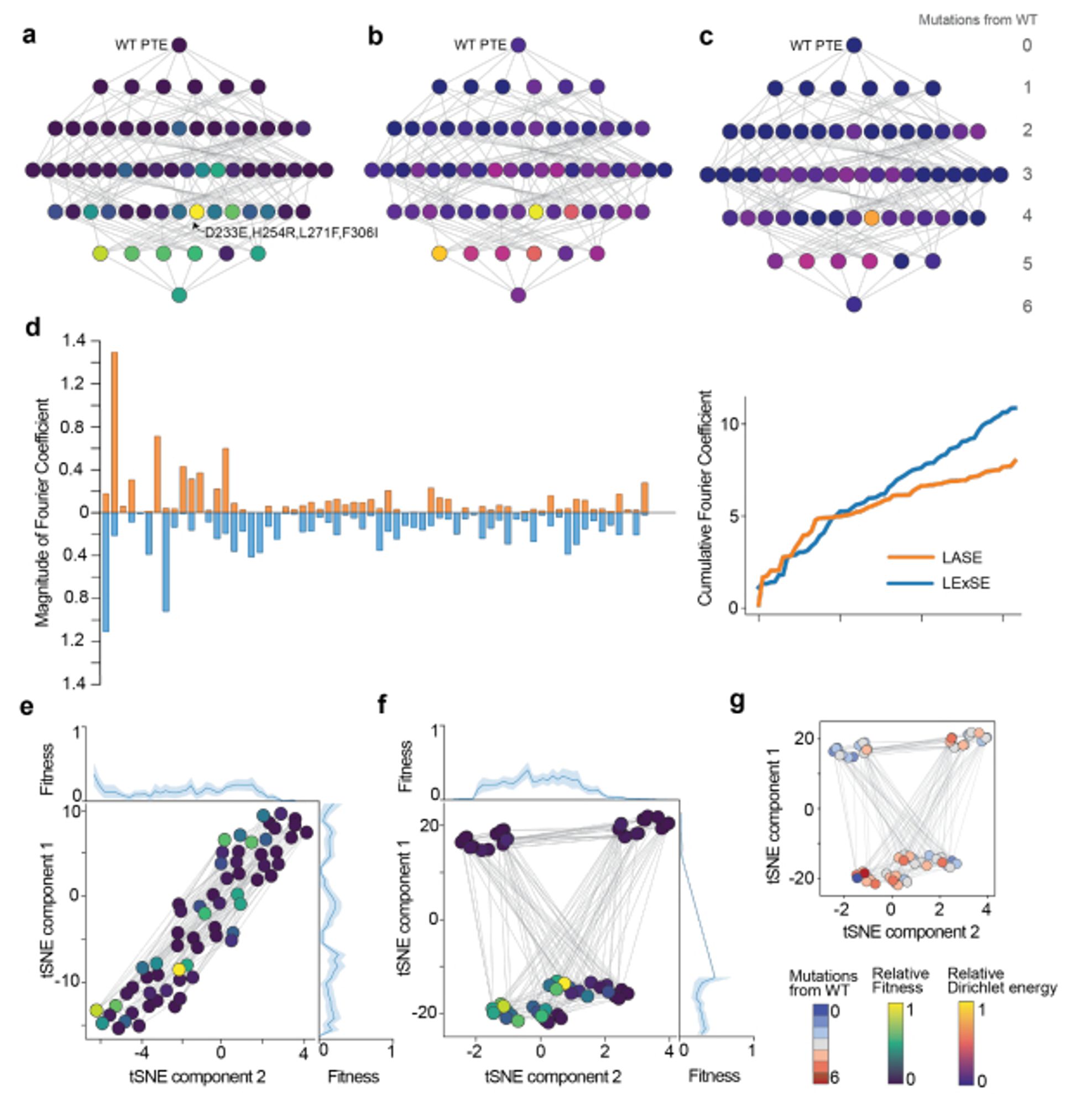
Use sequences from ancestral sequence reconstruction to train better local protein sequence representations.

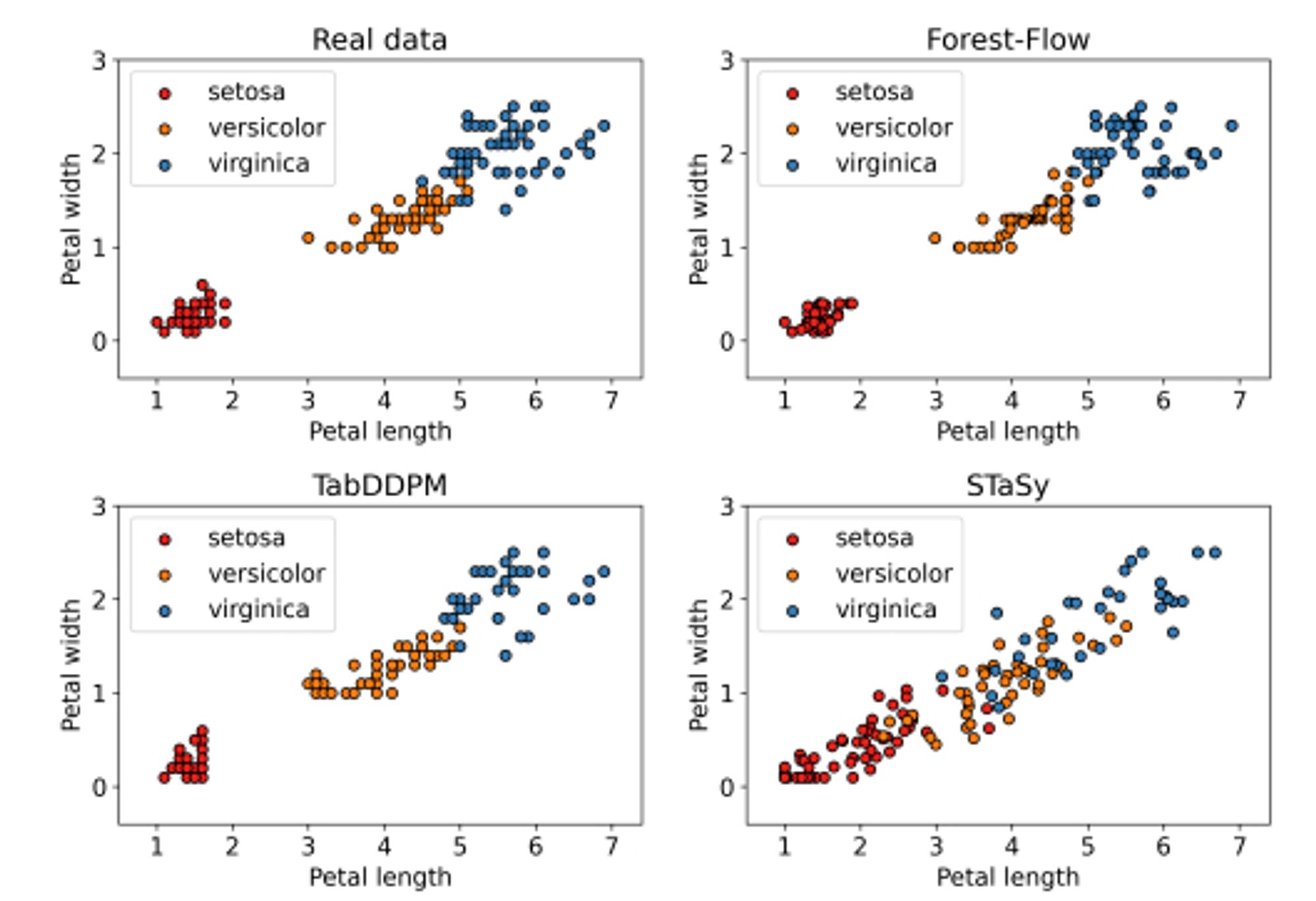
Diffusion and flow matching generative models implemented with XGBoost instead of neural networks! arxiv.org/abs/2309.09968
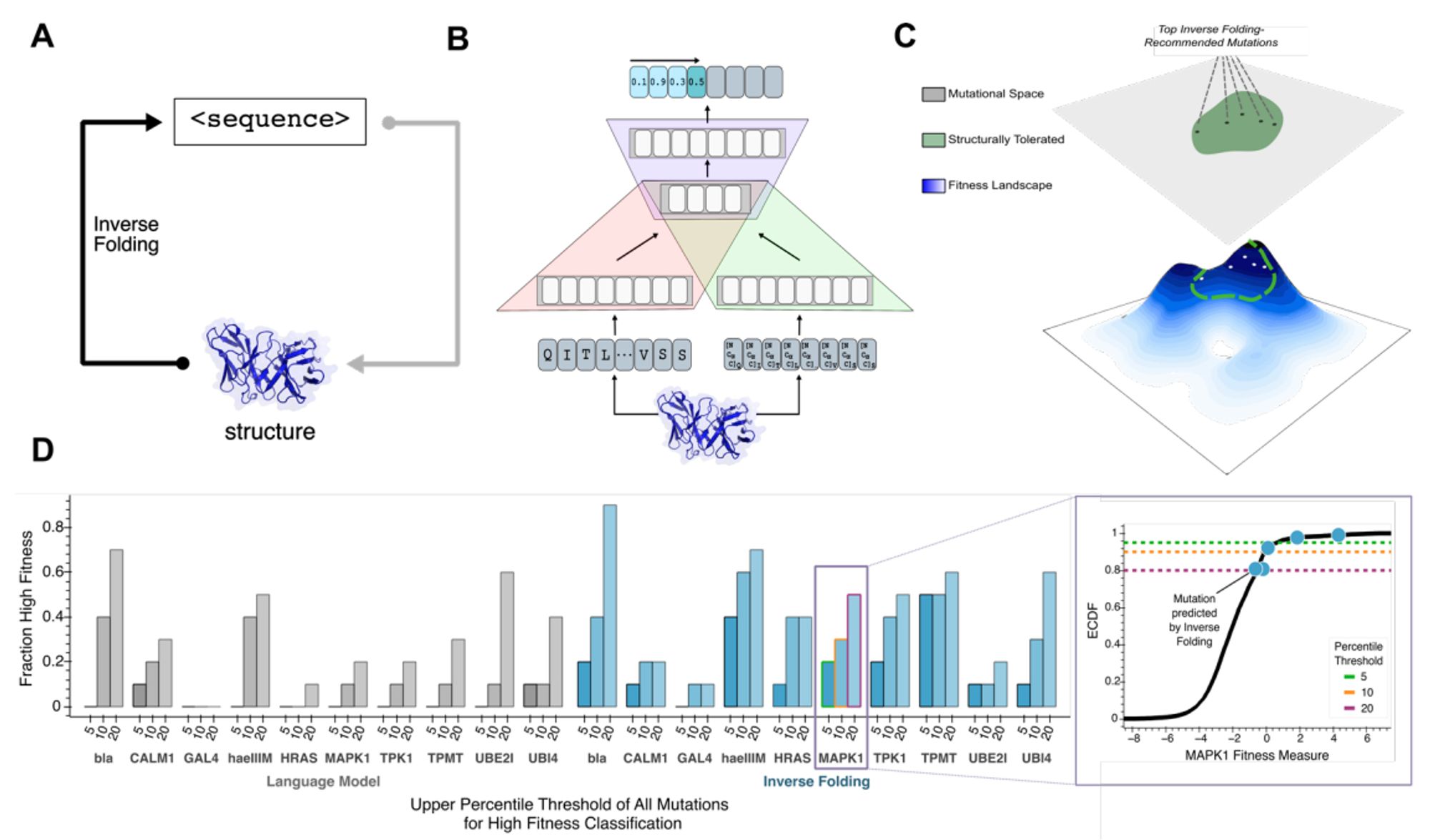
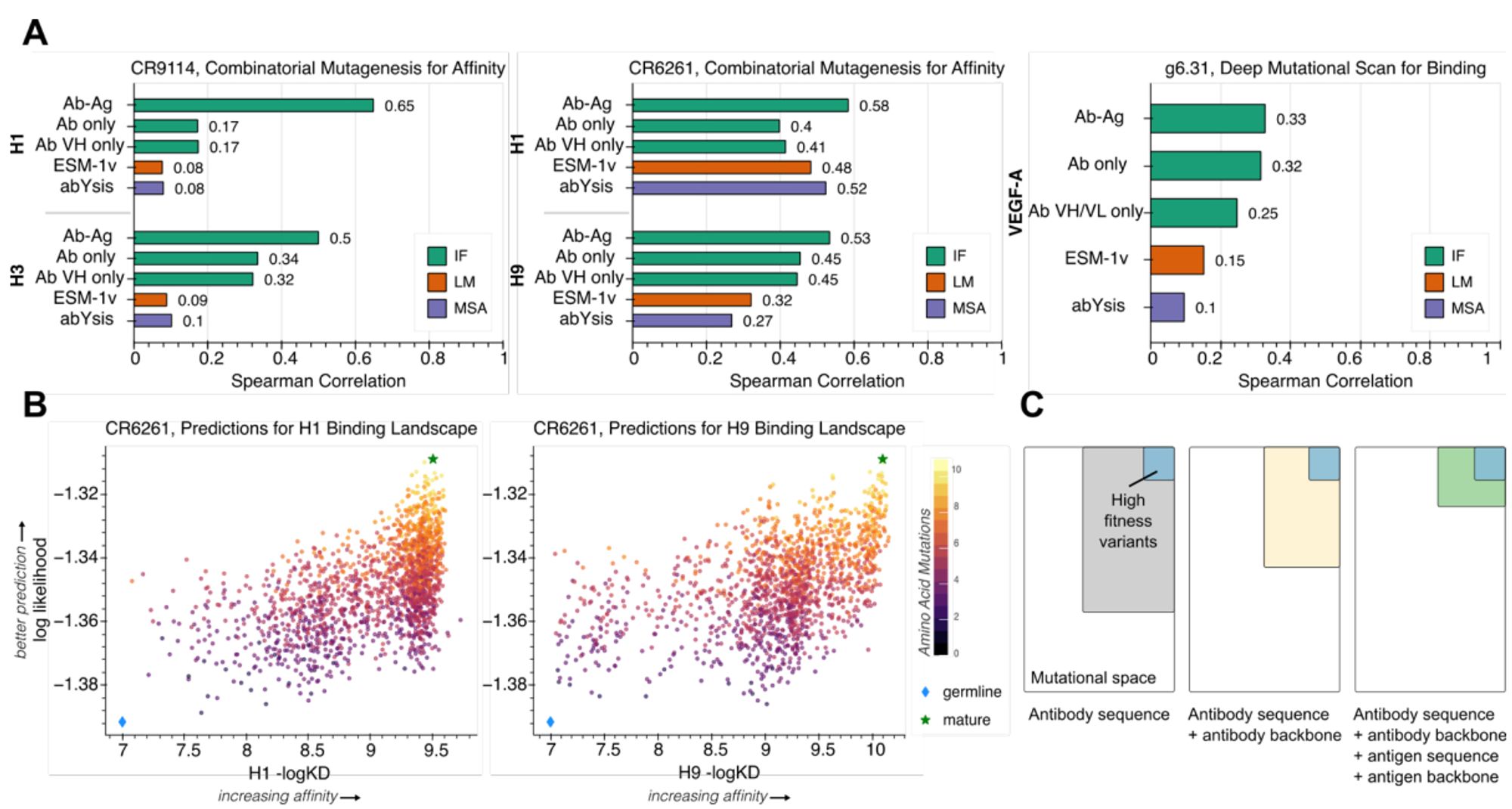
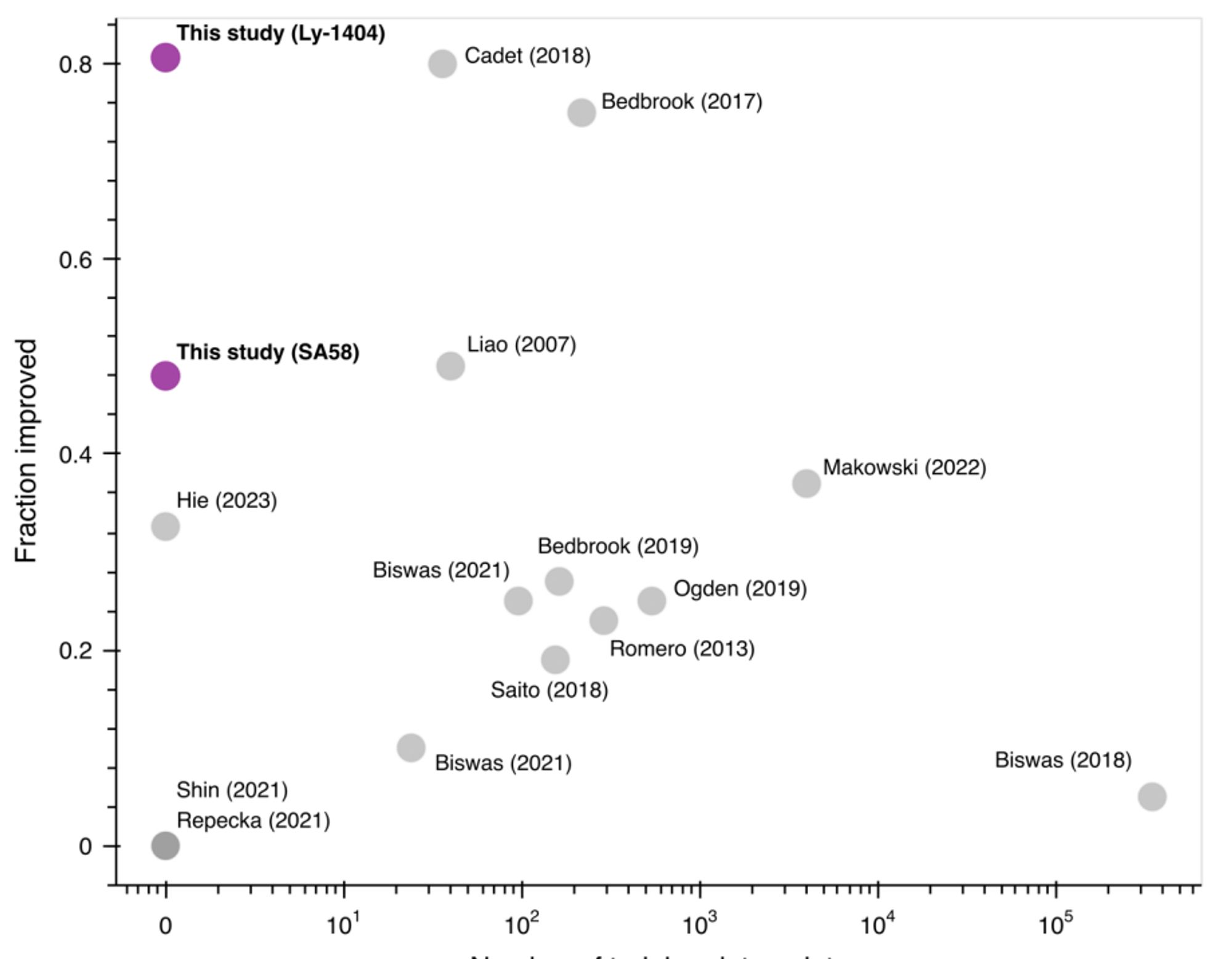
Use inverse folding likelihoods as an unsupervised protein sequence optimizer. www.biorxiv.org/content/10.1...
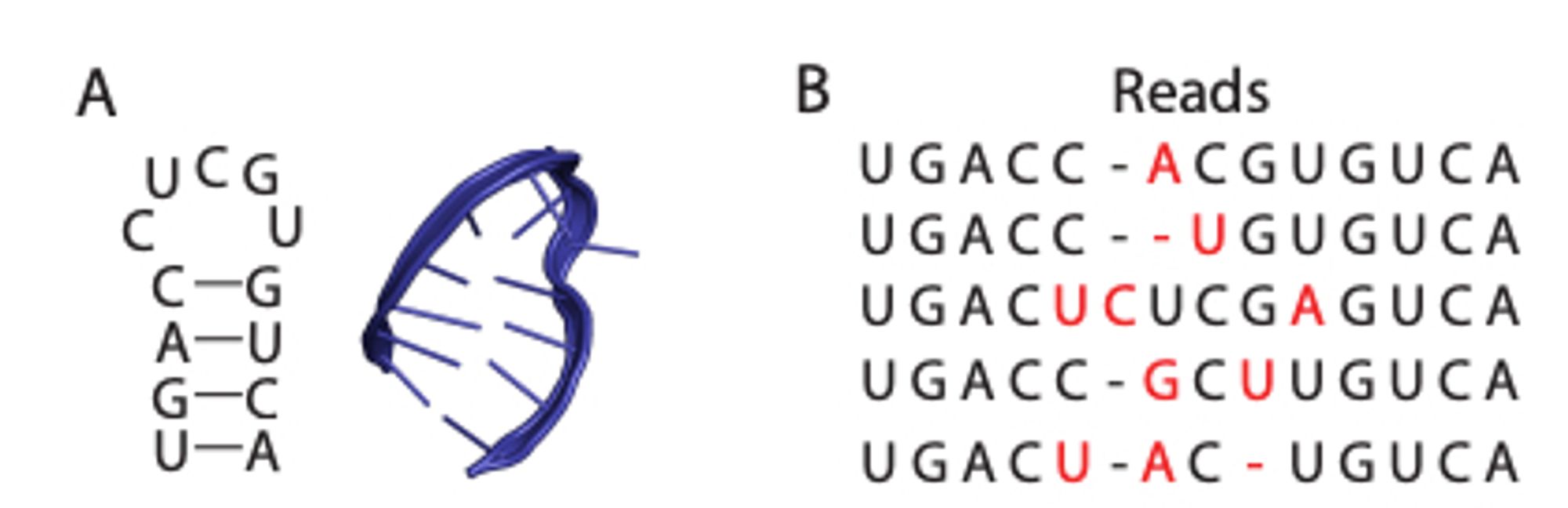
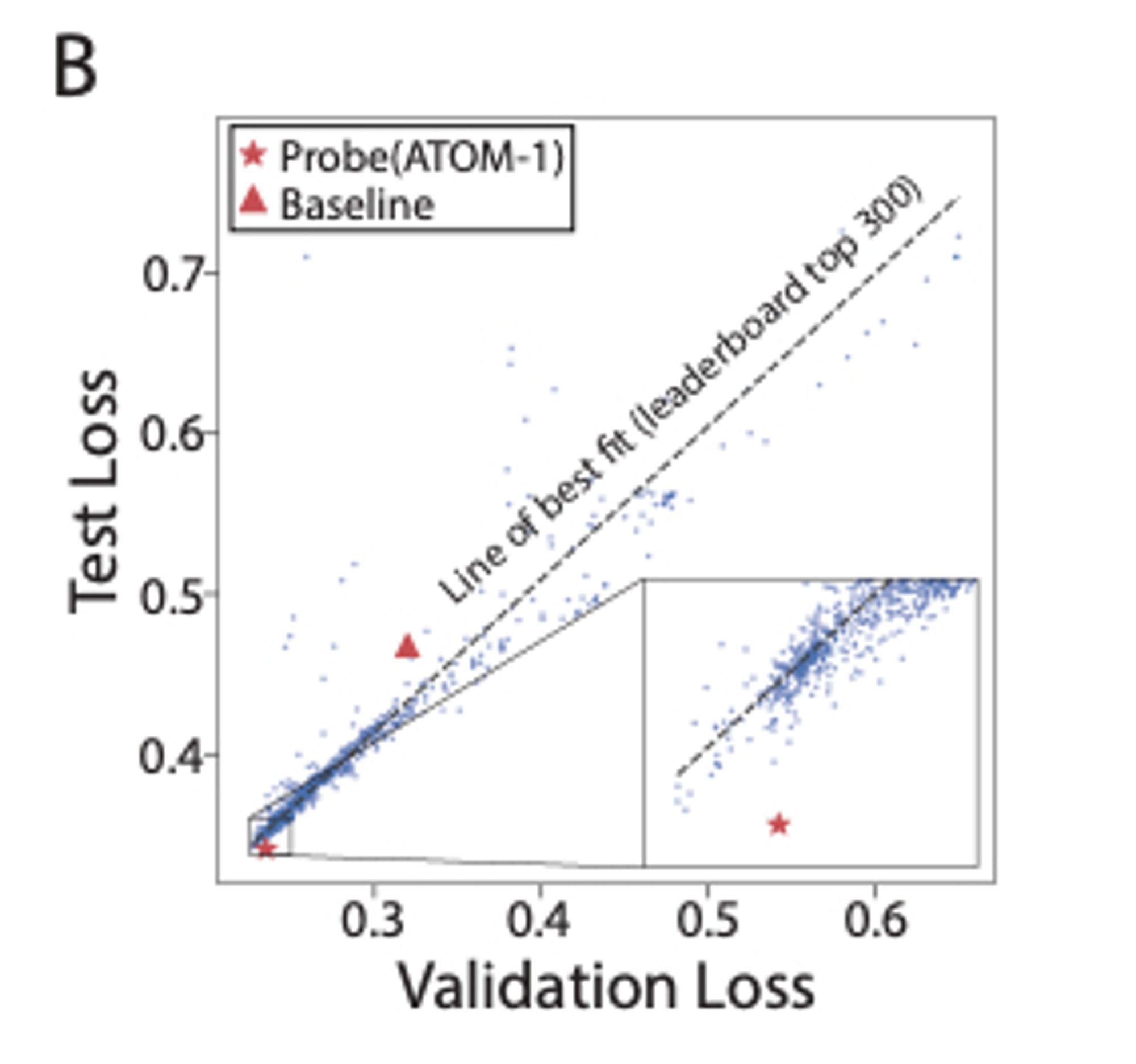
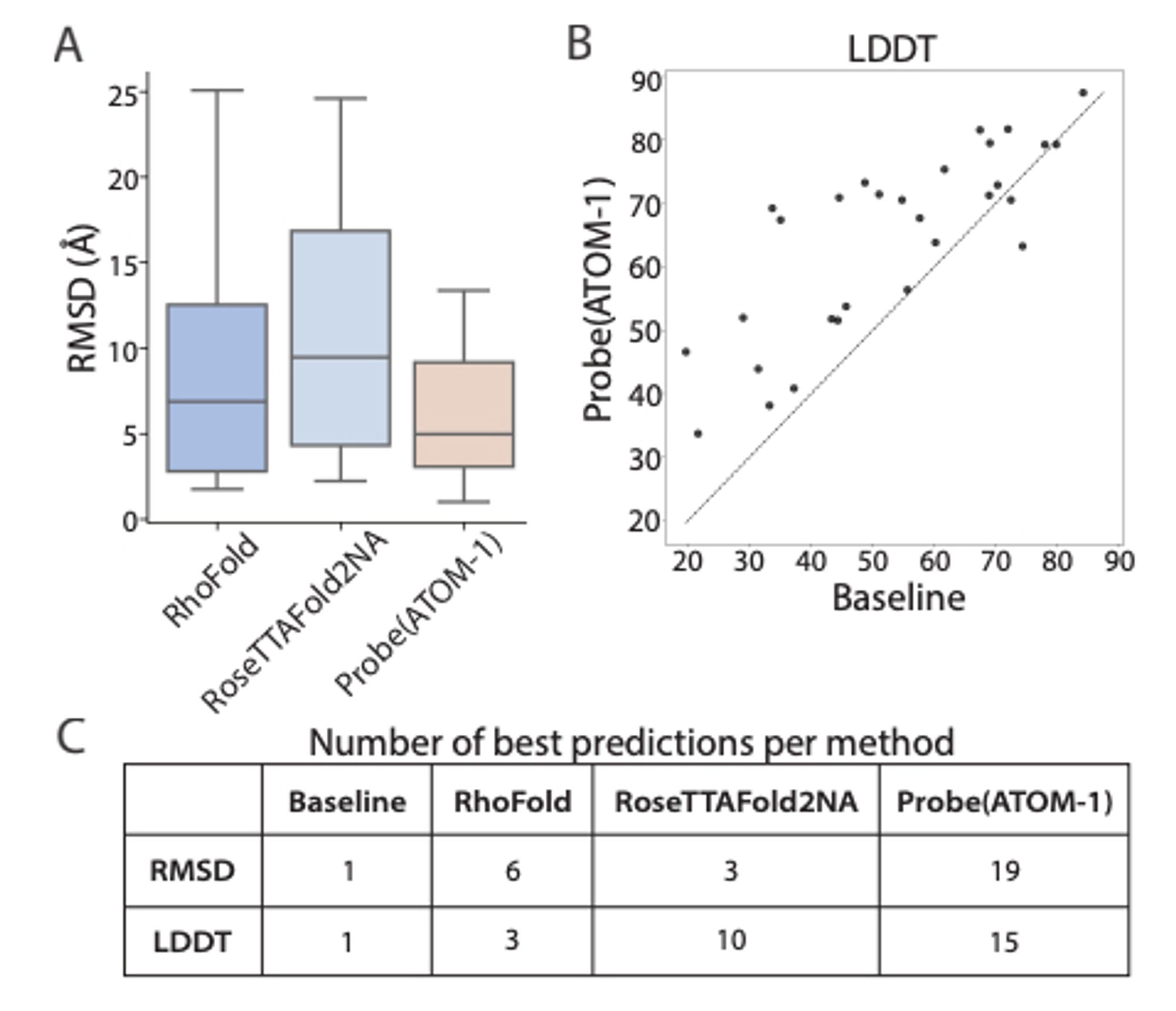
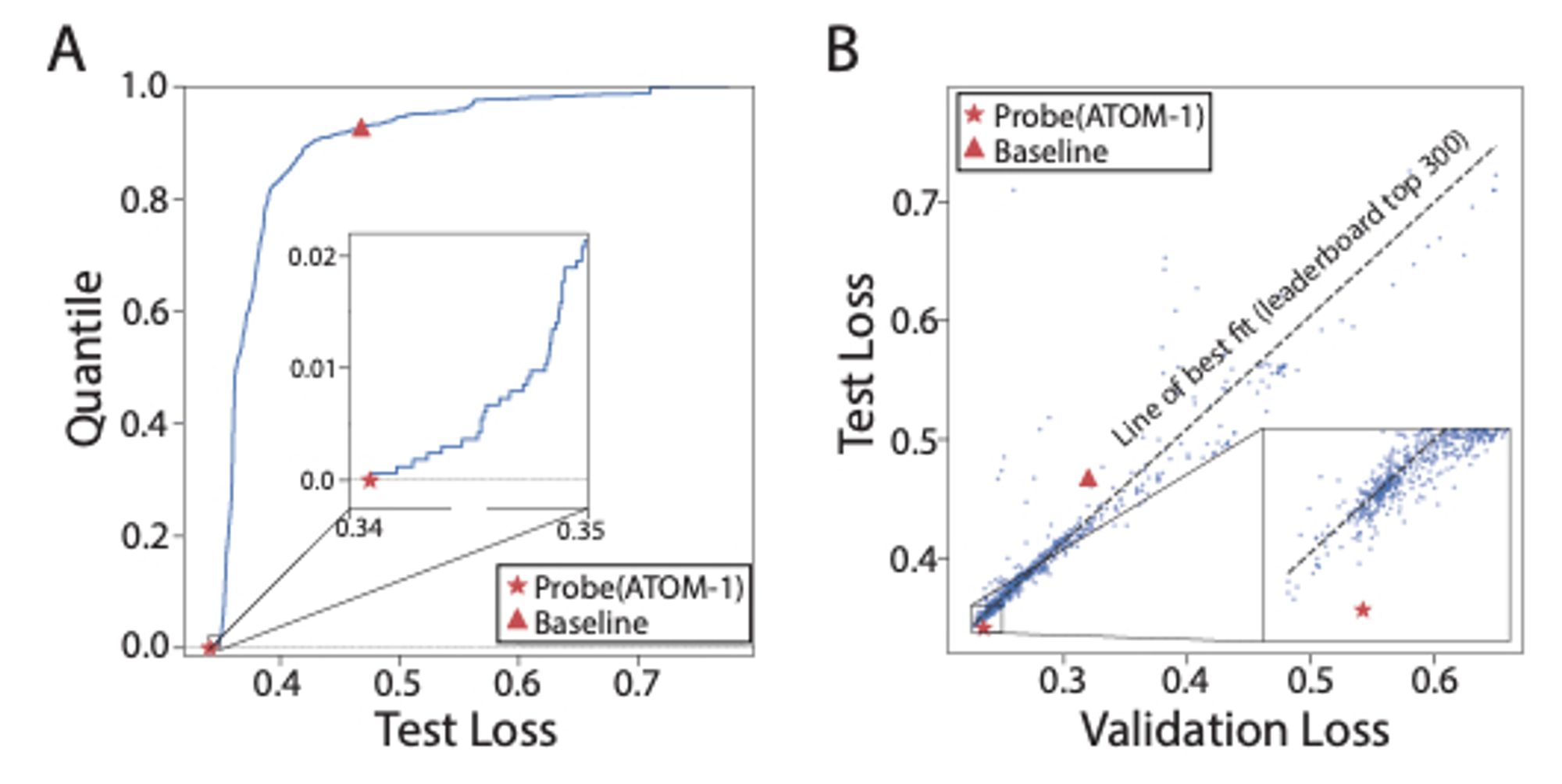
ATOM-1: An RNA foundation model trained to predict the results of chemical modifications to RNA sequences can be easily probed to predict RNA structure and properties. www.biorxiv.org/content/10.1...