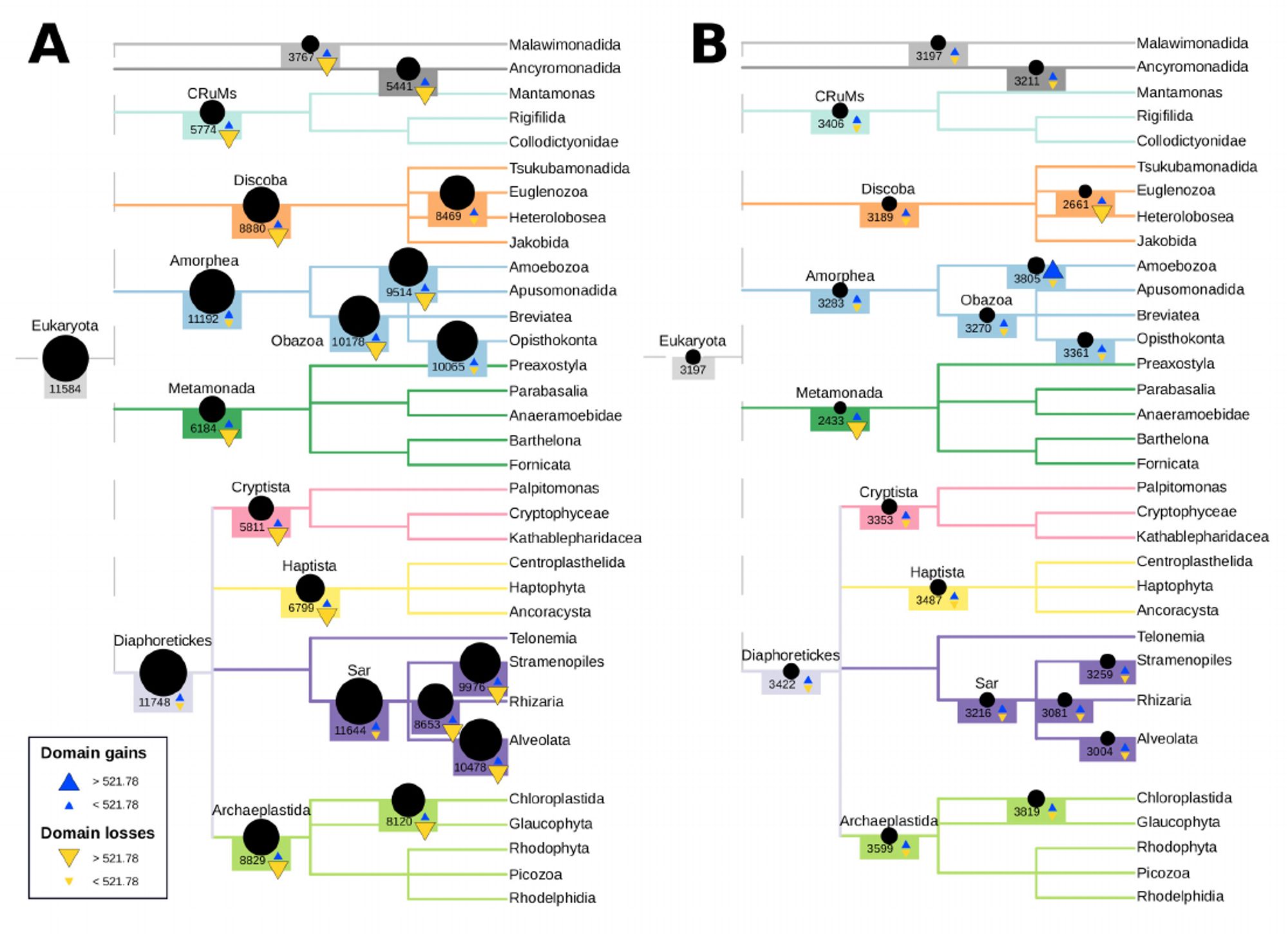
Next, we compared Dollo (A) to ML (B) on a real dataset of Pfam domains. Dollo’s LECA has more domains than any extant eukaryote followed by widespread losses; ML’s has fewer LECA domains and balanced gains/losses. Which to trust? Our results suggest caution with Dollo parsimony.
With Paschalis Natsidis and Max Telford’s simulated sets of 5,000 orthologs with no gains or losses, we tested our suspicion together. We compared Dollo to maximum likelihood (with Laurent Guégen), finding that Dollo consistently overestimates ancestral gene content towards the root.

New preprint! For years, we had worried that Dollo parsimony, which is based on principles developed for morphological characters, was not an appropriate choice for ancestral gene content reconstruction based on sequence homology. Now we can say: it’s not! doi.org/10.1101/2023...#protistsonsky

bioRxiv - the preprint server for biology, operated by Cold Spring Harbor Laboratory, a research and educational institution
The BEAP lab would like to welcome two new members! Xènia Maya joins us as a lab technician for the IBE Microbe Hub (joint with the Multicellgenome and @fonamental.bsky.social labs), and Viktoriya Petlovana joins us as a visiting scientist from Taras Shevchenko National University of Kyiv.


