likewise there’s no web version because if I were to make one it would just be a worse version of @deck.blue
Scientific writing tip: Don't remove content to fit a page or word limit. Remove words and phrases. You'll be surprised at how much you can cut, and your writing will probably get more interesting to read as a side effect.
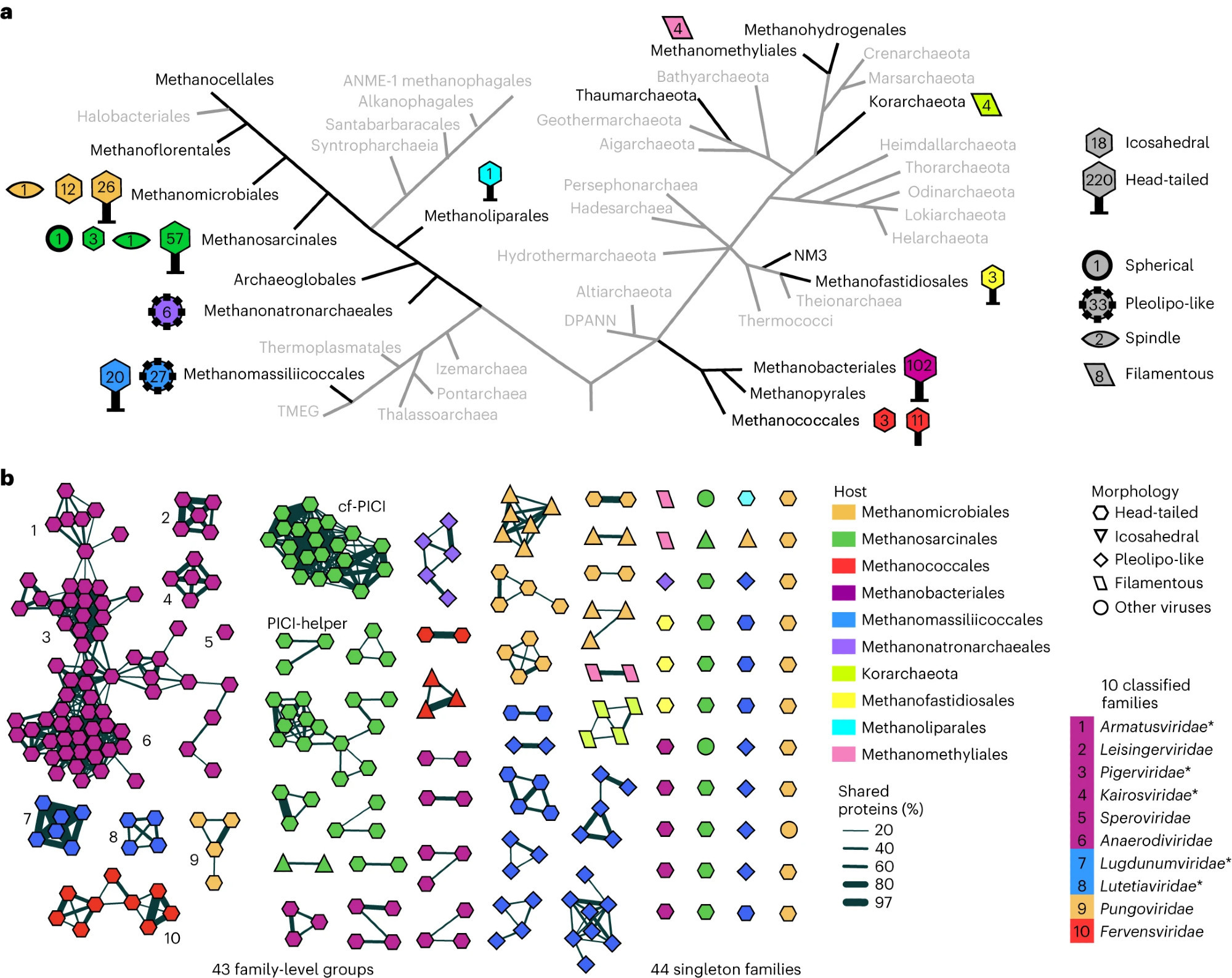
A compendium of viruses from methanogenic #archaea reveals their diversity and adaptations to the gut environment - a fruitful collaboration with colleagues from Institut Pasteur, Sofia Medvedeva, Simonetta Gribaldo and Guillaume Borrel. www.nature.com/articles/s41...
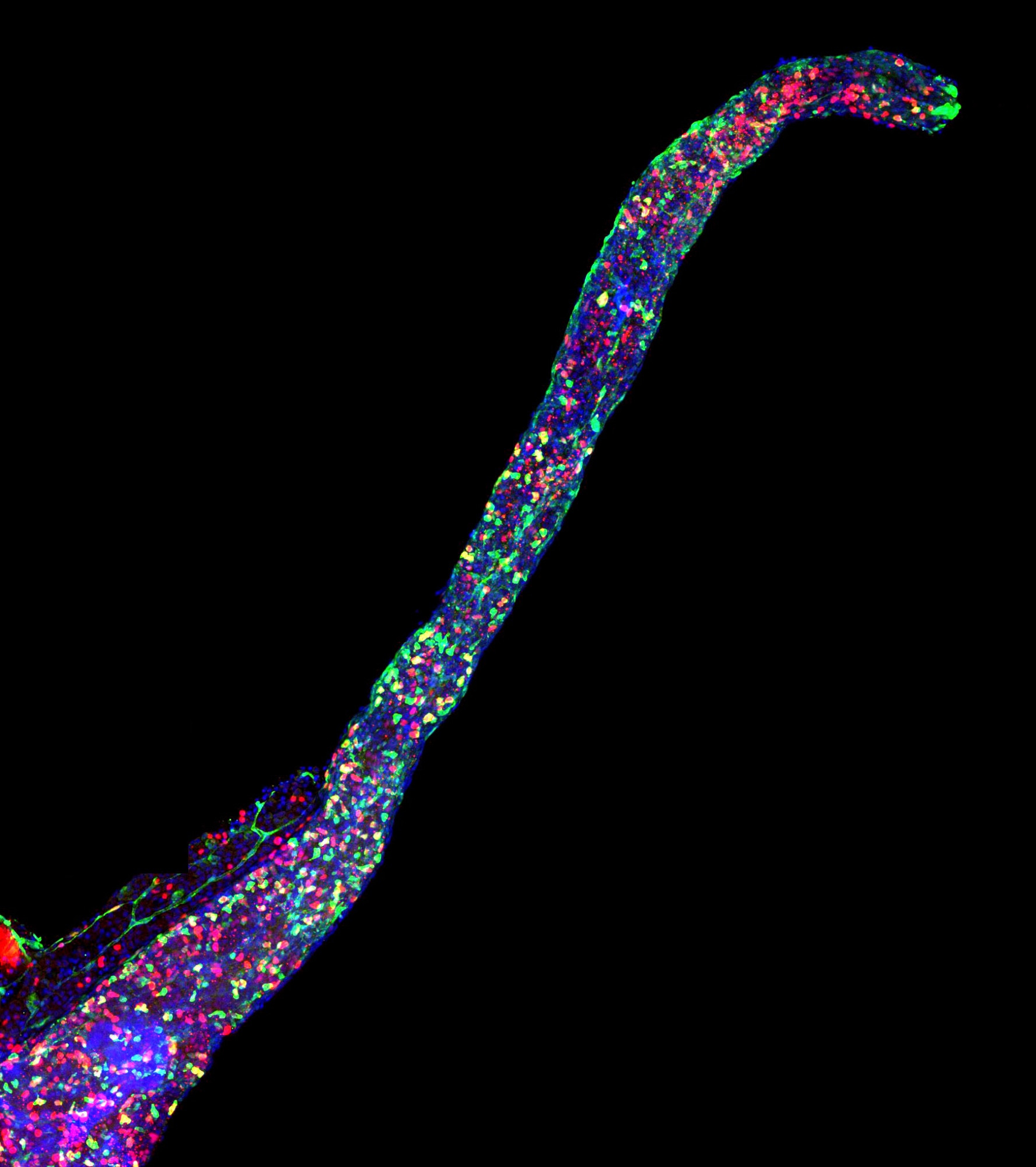
My lab's first big research paper is now out in mBio! 🎉 We ask: how can Legionella antagonize other microbes with reactive oxygen species without hurting themselves? Answer: several dynamics seem to generate a 'toxic moat' of ROS around producers pubmed.ncbi.nlm.nih.gov/37728338/
Awesome to see so many new arrivals! Reposting this to orient the microbiologists: The MicroSky feed is here: bsky.app/profile/did:...forms.gle/kiPC8vDKGDZV...bsky.app/profile/did:... Enjoy!
Sweet, i was wondering if there was a way to see people on the microsky feed list, to find people to follow. Thanks James!
Use the feeds. What's Science🧪: bsky.app/profile/did:...bsky.app/profile/did:...bsky.app/profile/did:...
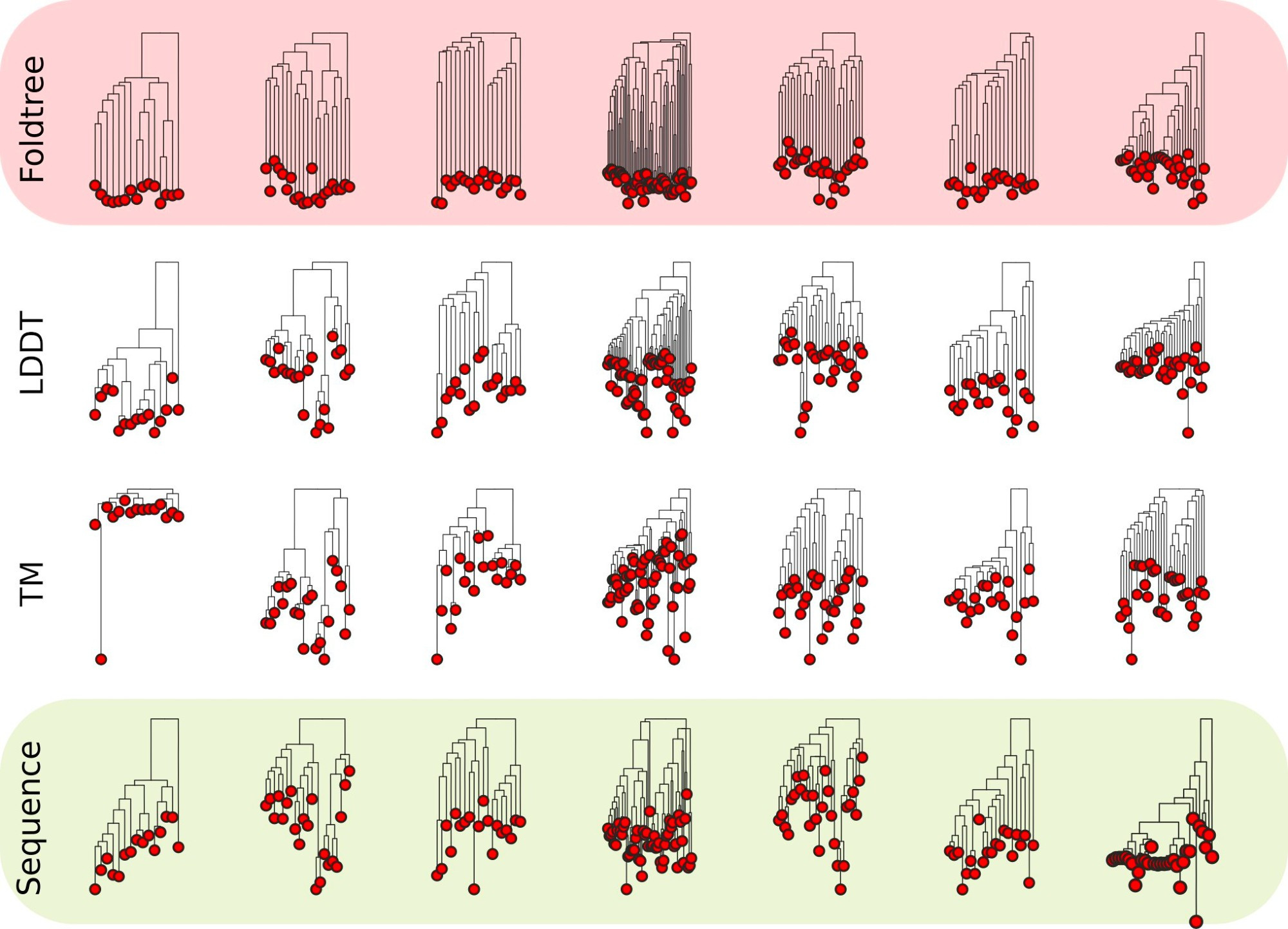
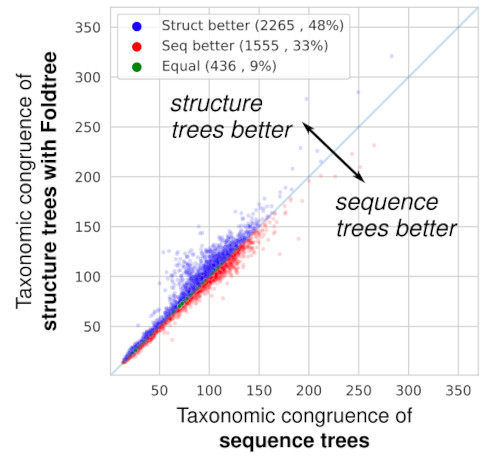
A lack of reliable structural protein predictions has hindered benchmarking structure-based phylogenetic inferences compared to sequence-only models. Here Moi et al., describe FoldTree showing that structure-based trees can outperform sequence-based trees (sometimes). www.biorxiv.org/content/10.1...