@dsmzd3.bsky.social Welcome to MicroSky!
Happy to share the final version of our paper on recombination & gene flow in the human gut microbiome. Heroic effort by @zzzhiru.bsky.social getting it over the finish line (>40 SI figures!) If you're interested the interplay between recombination & selection in the gut, Zhiru is the one to ask!
Excited (and relieved!) to share that my 1st paper in grad school finally came out in PLOS Biology! We were motivated by a very basic question: how fast, really, do bacteria exchange DNA in the human gut? (1/n)
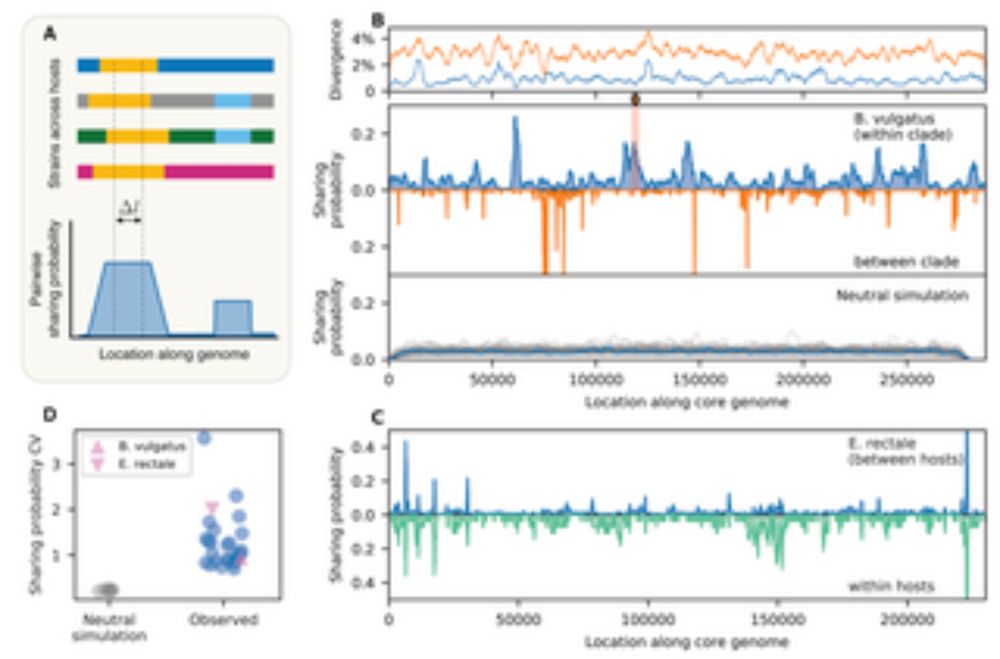
Recombination is a ubiquitous force in bacterial evolution, but its dynamics are poorly characterized in many natural populations. This study presents a metagenomic approach for quantifying the landsc...
A good review of long-read metagenomics, but not comprehensive. It is hard to stay current - the field is moving fast! Several tools appeared in the last 6 months: - assembly (metaMDBG) - binning (SemiBin2) - profilers/classifiers (Metabuli, Taxor, sylph, MetageNN, Melon) doi.org/10.1186/s129...
I've just added you to the Microsky contributors list - your post should appear momentarily
The Evolutionary Genomics of Complex Traits lab led by Luisa Pallares is looking for a postdoc to work on the genetic basis & evolution of phenotypic robustness of complex traits. The position is funded for 3 years. Deadline to apply 29.2.24. More info:tinyurl.com/36ee8tw4
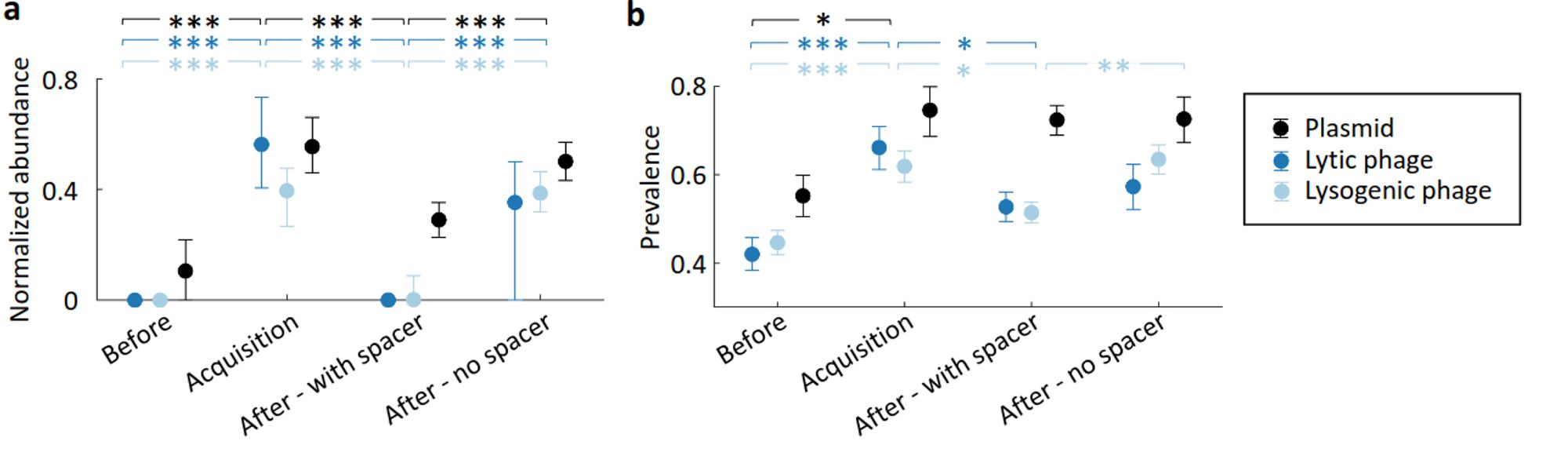
This is a nice longitudinal metagenomic analysis of CRISPR dynamics in the gut with a lot of cool findings, including showing clearance of MGEs after spacer acquisition: Dynamics of CRISPR-mediated virus-host interactions in the human gut microbiome www.biorxiv.org/content/10.1...
Comparative analysis of metagenomic classifiers for long-read sequencing datasets bmcbioinformatics.biomedcentral.com/articles/10....github.com/lbcb-sci/Met...
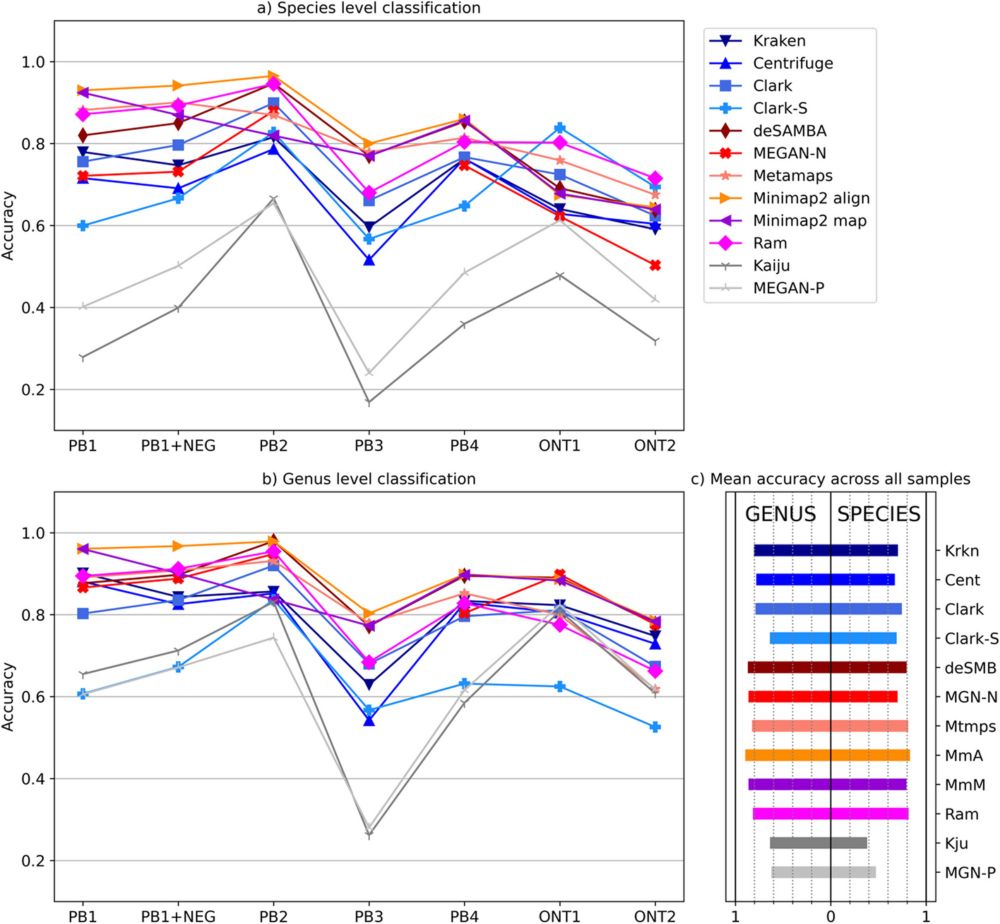
Background Long reads have gained popularity in the analysis of metagenomics data. Therefore, we comprehensively assessed metagenomics classification tools on the species taxonomic level. We analysed ...
MicrobeMod 1.0.3 is released! It is now compatible with the dual 4mC and 5mC experimental all-context base calling model. 4mC calling seems to be working well for me so far, but want to see more tests! Excited to get all 3 major prokaryotic methylation types now. github.com/cultivarium/...

A toolkit for exploring prokaryotic methylation and base modifications in nanopore sequencing - GitHub - cultivarium/MicrobeMod: A toolkit for exploring prokaryotic methylation and base modificati...
An excellent resource for microbiome-related conferences in 2024. I've pinned this to the top of the #MicrobiomeSkybsky.app/profile/did:...