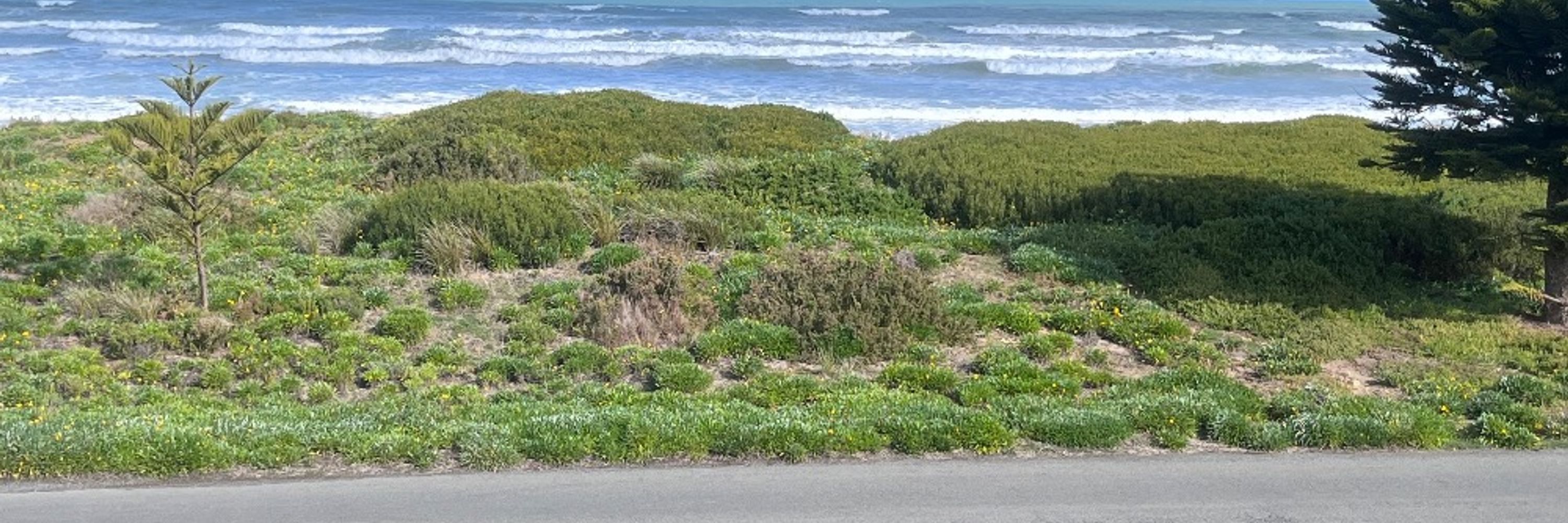

TARGet will utilise recent genomic advances to better understand AMR and leverage this information for surveillance, diagnostic, and infection prevention control practices. It has led jointly by @ksbakes.bsky.social (U Cambridge) and myself (U Birmingham). /2
Thanks to everyone involved including both old and new collaborators @micro_deb @BenjaminHowden @DrDanielleIngle @MarieChattaway @rodwell_ella @neilhall_uk and, as always, amazing teamwork with @MalakaDeS @ChalotteChong_ and @RJBengtsson 8/8 (with apologies to those who switched handles from X)
So, welcome to your new genomic era understanding of an old disease threat: S. Panama! It was an honour co-supervising @1caiseypulford’s PhD work with @jay_salsa and working with the tireless @weill_xavier who absolutely made this work happen 7/8
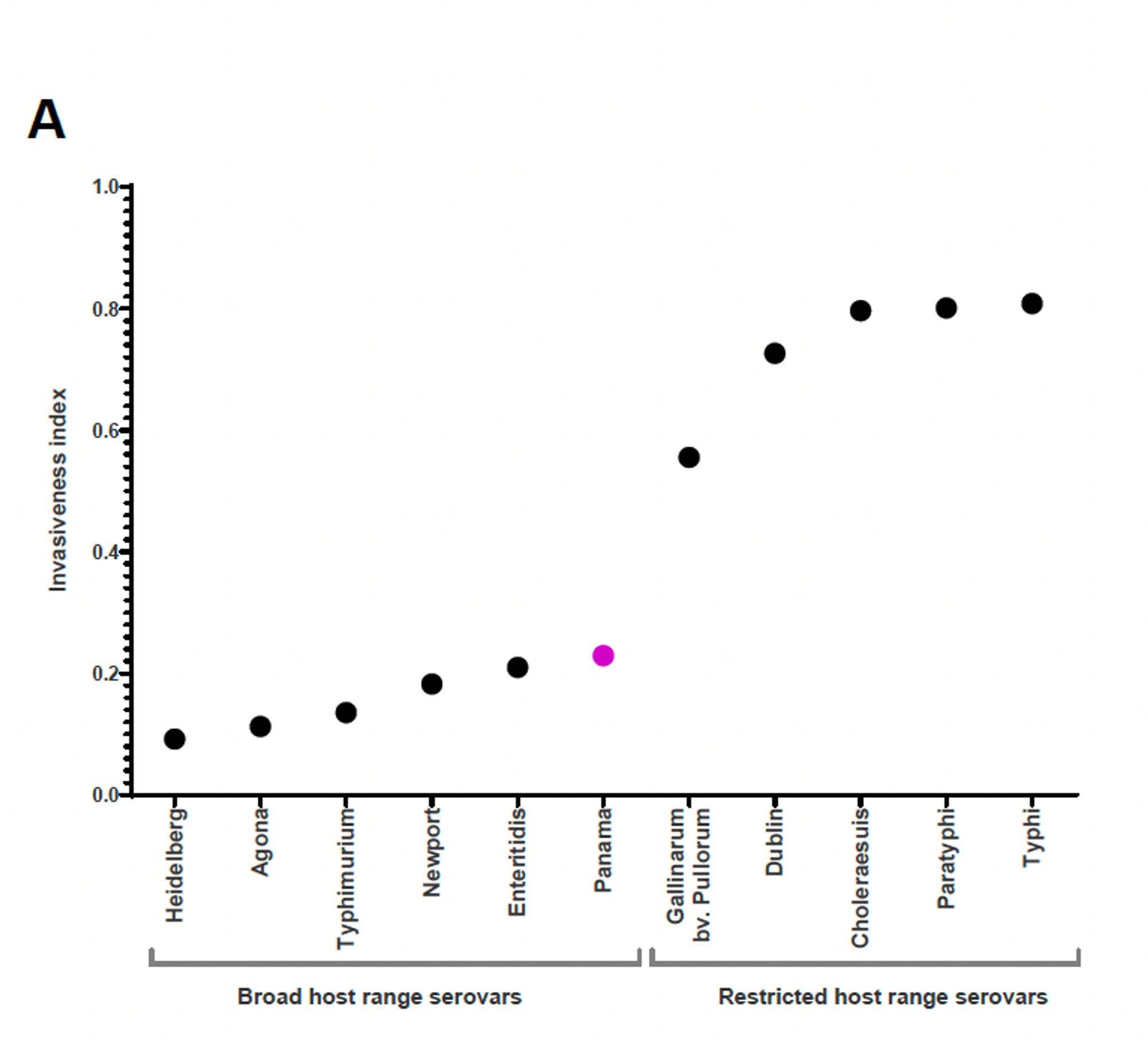
The ‘invasiveness index’ (thanks @nwheeler443) of S. Panama was elevated relative to other broad host-range Salmonella serovars and highest in the European- AMR- associated Clade 2, suggesting this might pose the greatest public health threat 6/8
Predicted AMR was carried by gene cassettes against some old favourites (streptomycin, ampicillin, chloramphenicol, trimpethoprim, sulfisoxazole, and tetracycline) and one XDR isolate with further fluoroquinolone, polymyxin, and 3rd generation cephalosporin resistance 5/8

There were low rates (14.5% of isolates) of AMR predicted in S. Panama and AMR was typically found in the clades associated with Europe and Asia 4/8
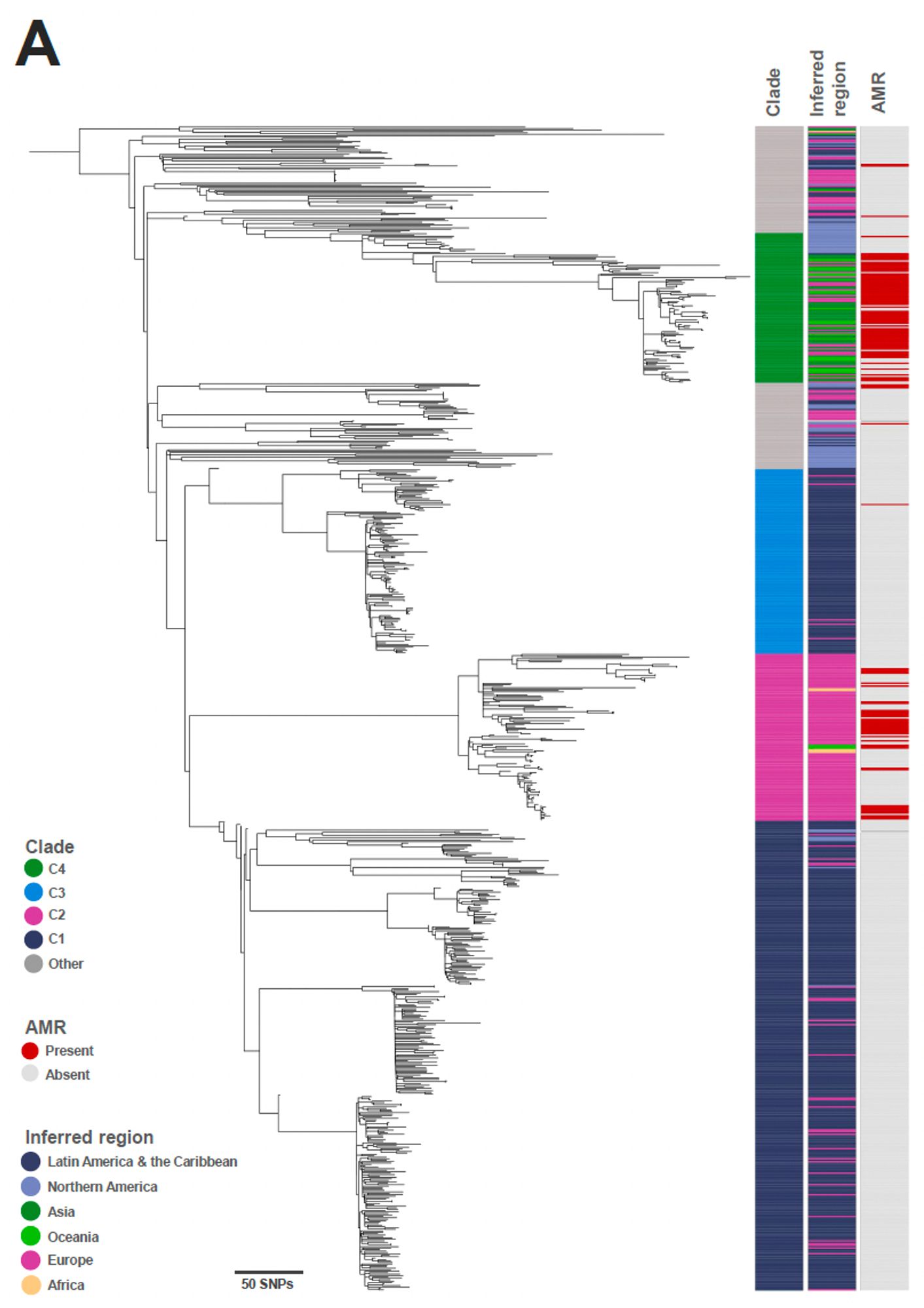
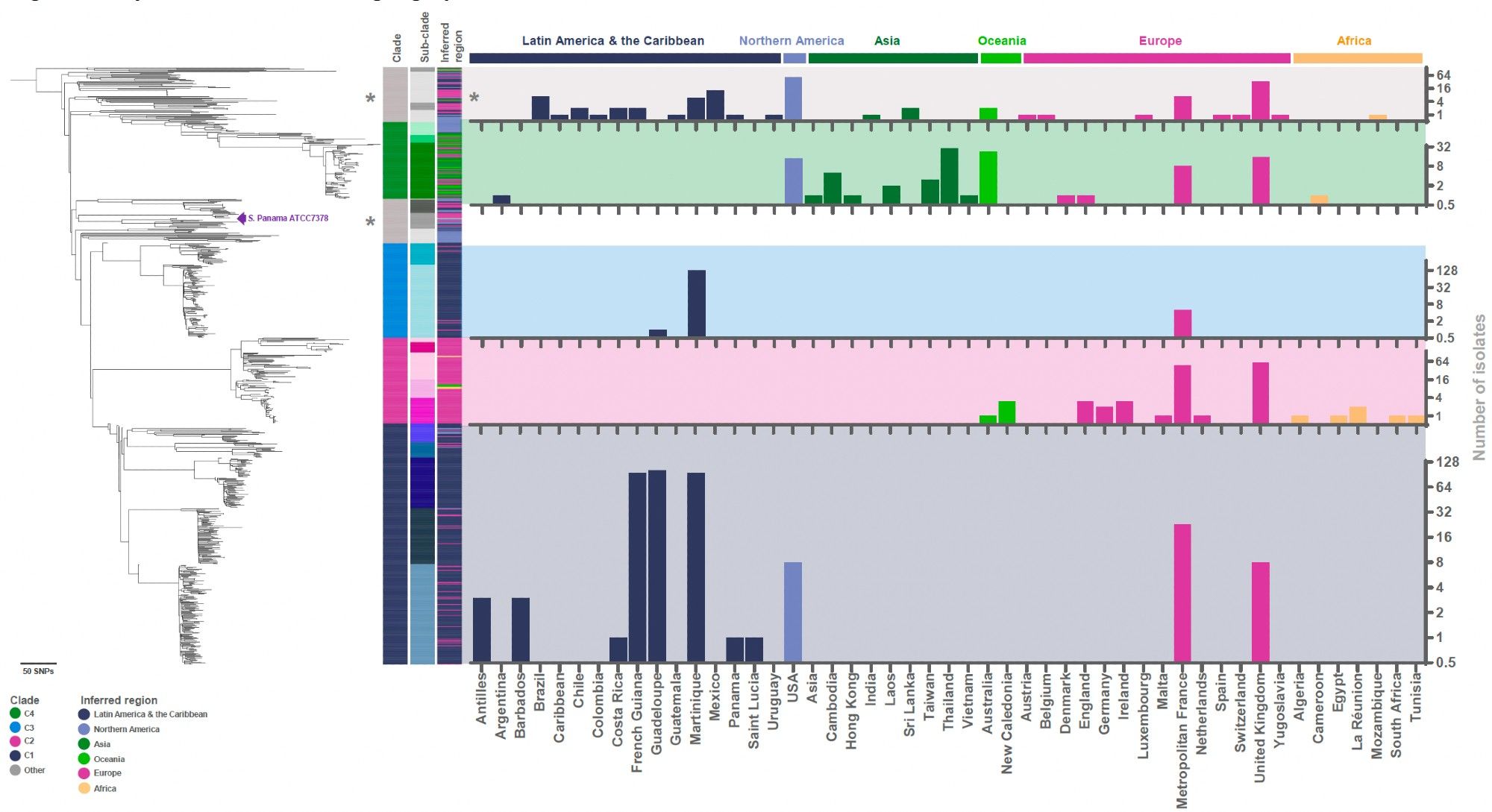
The usual tricks (phylogeny and BAPS clustering) revealed four geographically associated clades, including some further island-associated substructure in the Caribbean 3/8
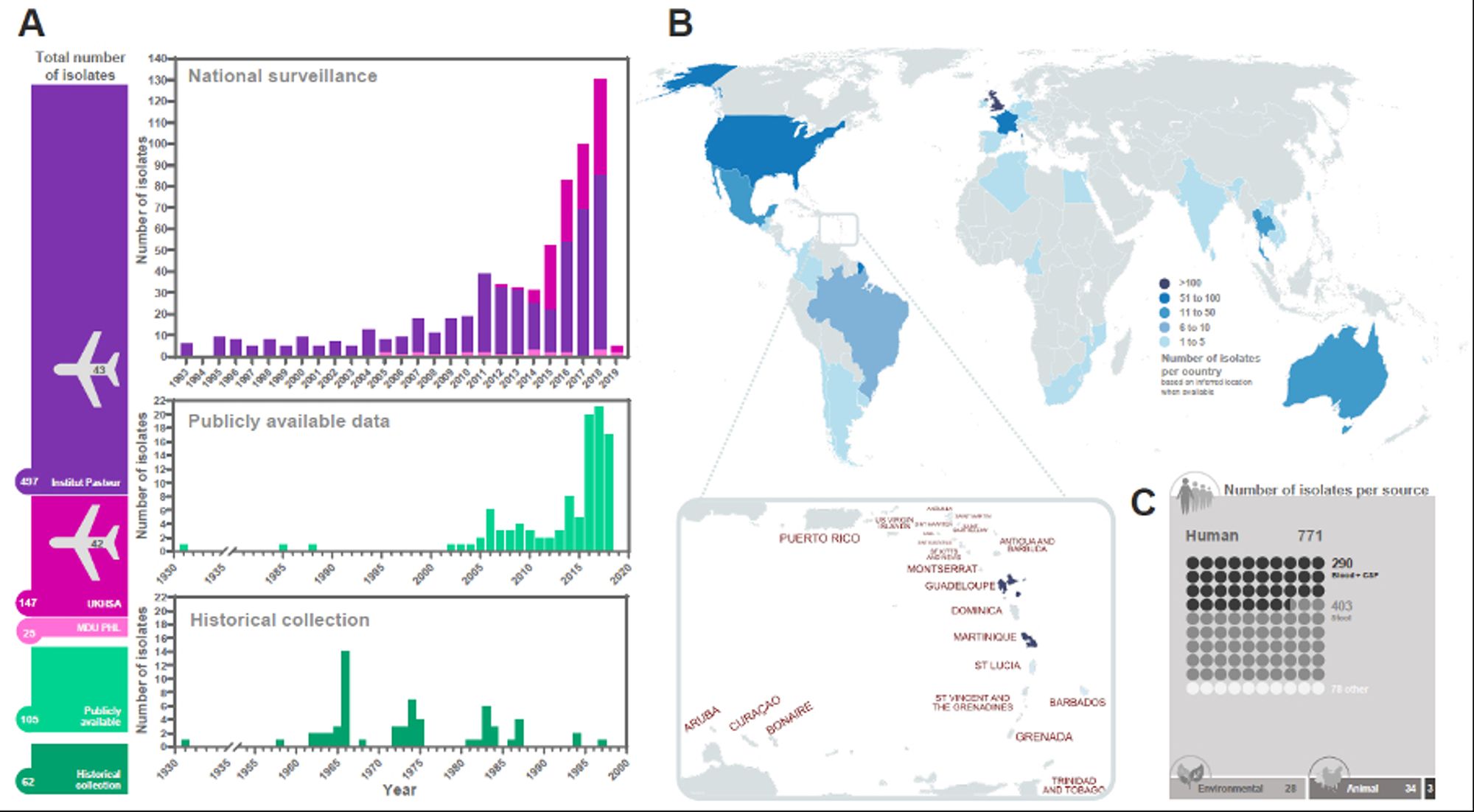
Together with an amazing set of public health collaborators we assembled a diverse set of 836 S. Panama genomes from different disease presentations and global regions spanning nearly 100 years 2/8
No thanks Lars - all set
