For background on this unhealthy obsession with linking taxonomic names to their original descriptions see doi.org/10.3897/BDJ....

A major gap in the biodiversity knowledge graph is a connection between taxonomic names and the taxonomic literature. While both names and publications often have persistent identifiers (PIDs), such a...
I feel like this also demands a link to possibly the best website in the world: HORG are the go-to for any info on occlupanid morphology www.horg.com
HT to @phypapers.bsky.social for inspiration, although I’ve taken a different route to get EvolDir here, it helped nudge me to get this done.
Note that this is a crude bot (same one as used for Twitter and Mastodon). If you want to post an announcement on EvolDir you’ll need to do it via email, see link on @evoldir.bsky.social profile page.
GBIF portals won’t work because, as far as I know, they require the data to be in GBIF and queryable using their API. I don’t want to add data to GBIF, so much as add an annotation (the JSTOR barcode identifier).GBIF portals won’t work because, as far as I know, they require the data to be in GBIF and queryable using their API. I don’t want to add data to GBIF, so much as add an annotation (the JSTOR barcode identifier).
The discoverability of types in JSTOR Plants is easier than in GBIF, so a portal might help people find the equivalent records in GBIF more easily. It’s also still worrying that there are records in JSTOR Plants that are only available there.The discoverability of types in JSTOR Plants is easier than in GBIF, so a portal might help people find the equivalent records in GBIF more easily. It’s also still worrying that there are records in JSTOR Plants that are only available there.
Sad news indeed.Sad news indeed.
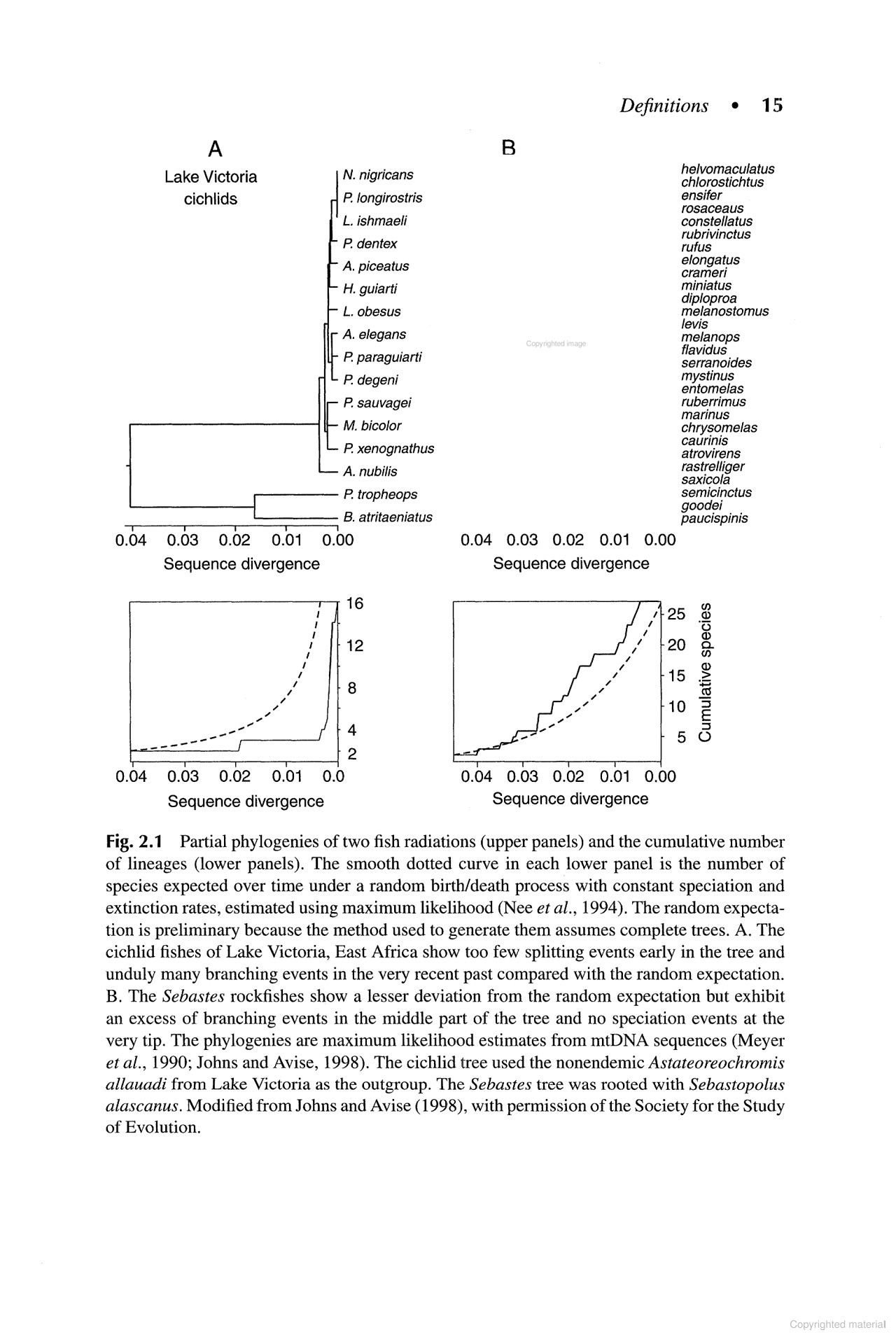
Found it! It's from "The Ecology of Adaptive Radiation” by Dolph Schluter, on page 15. You can see it in Google Books www.google.co.uk/books/editio...
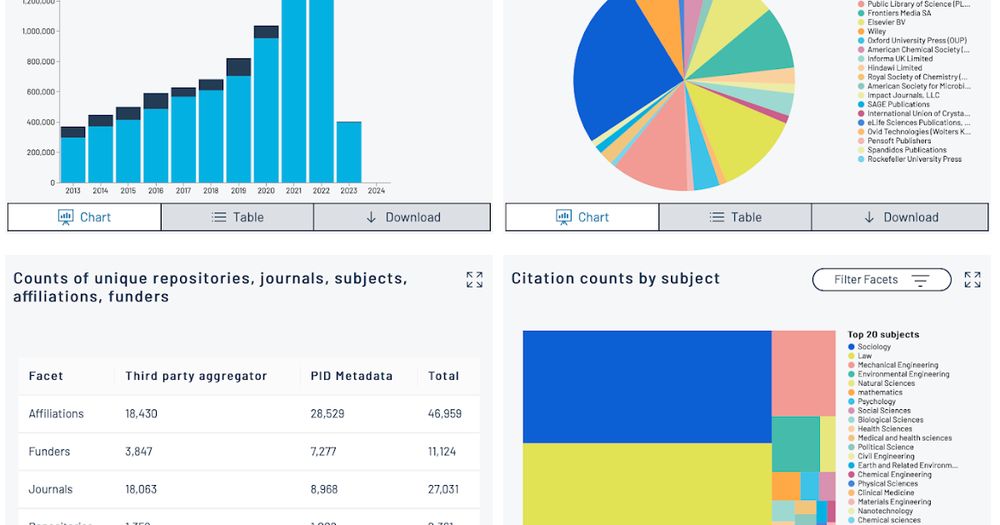
I wrote some comments on MakeDataCount here doi.org/10.59350/t80.... The single biggest category of data cited is GenBank sequences, which opens up the possibility of tracking the use of sequence data beyond their initial publication (e.g., reuse in phylogenetic studies).