If you’re not sold on scRNAseq, this is my favorite analogy on why measuring average properties of diverse samples can be misleading. What are the average properties of an animal at a zoo, and what would that actually tell you about that zoo? Bulk transcription of blood is kinda like this.
The preprint has detailed methods & metrics of equivalence; please dive in! We’d love your thoughts. This is only helpful if people use it! We think it lowers the barrier to embedding scRNAseq in large RCTs, and at underresourced settings (with resourced partners… scRNAseq is still $$$).
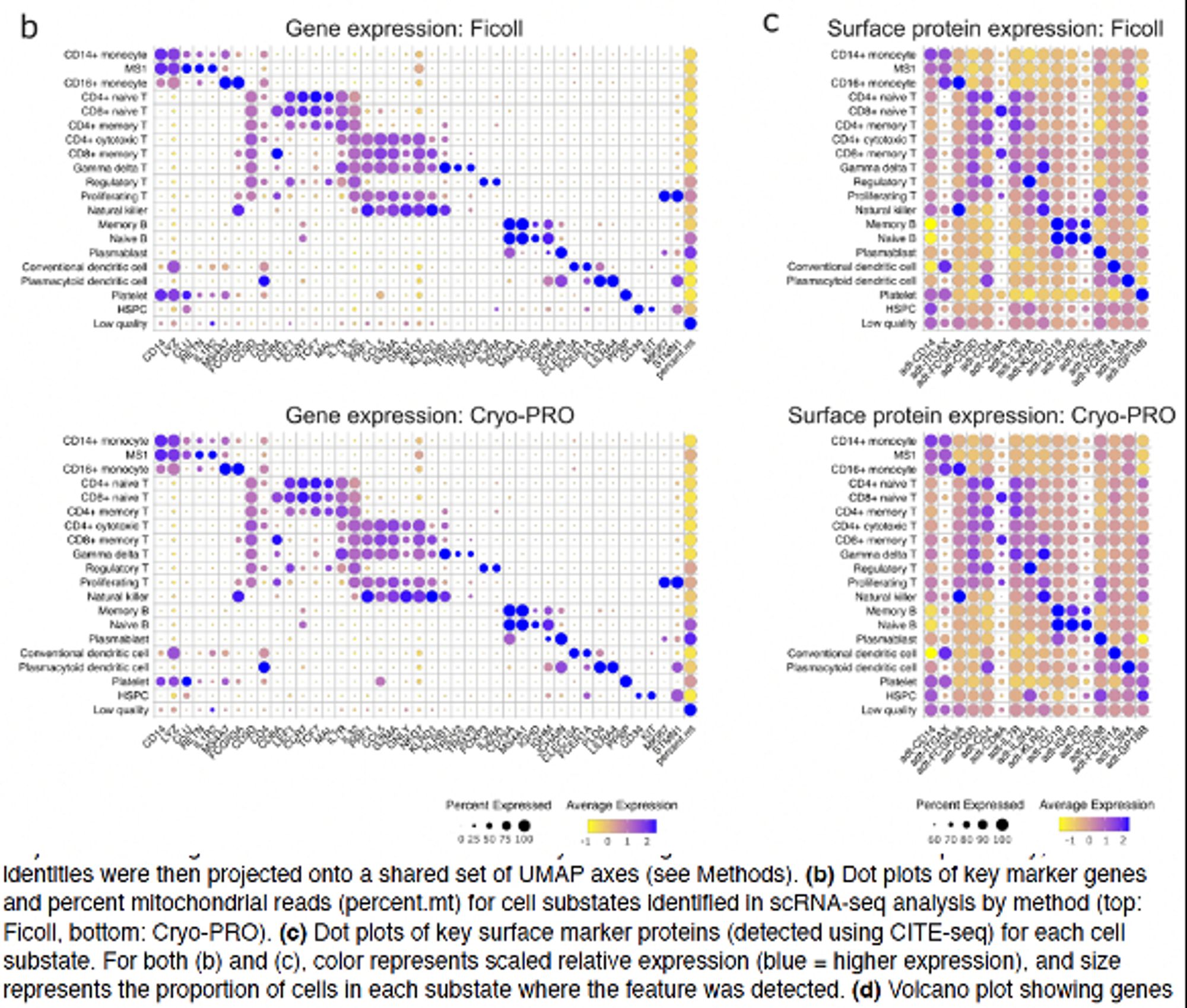
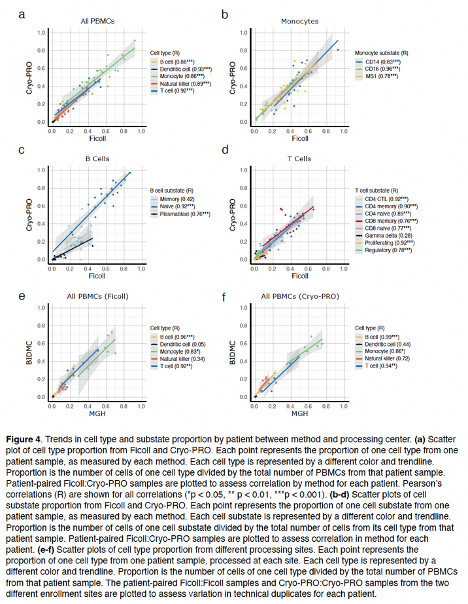
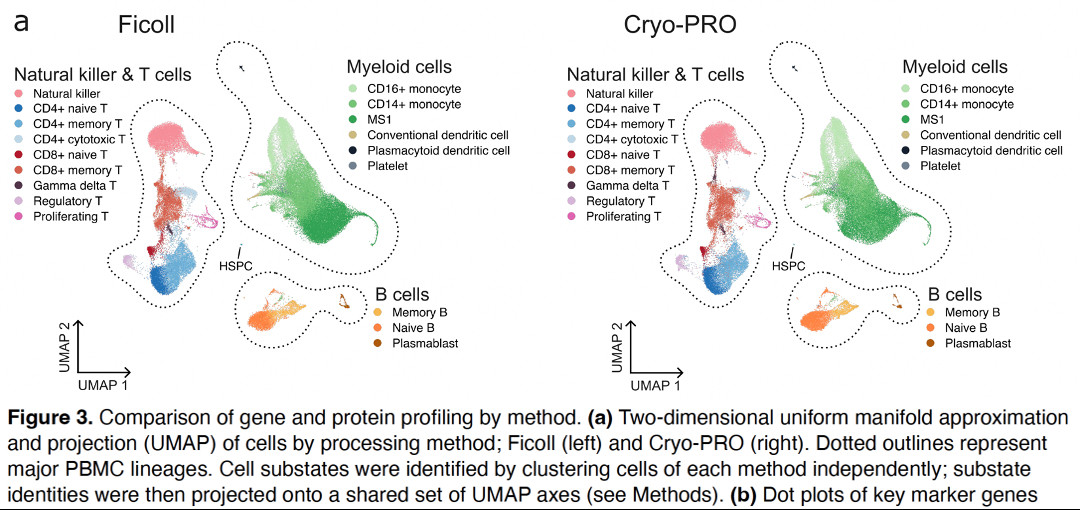
Beyond QC metrics, we independently clustered scRNAseq data obtained from 32 samples from 23 pts with our new method (“Cryo-PRO”: CRYOpreservation w Pbmc Recovery Offsite) vs Ficoll, then projected onto the same UMAP axes. Cryo-PRO data alone found all cell types & states, in comparable abundances.
Using the “old way” for scRNAseq, we previously found a new transcriptional substate of monocytes enriched in sepsis. But we could only recruit at high-resourced academic medical centers. The ability to do scRNAseq with minimal work will enable this method to be applied anywhere with a -80 freezer.
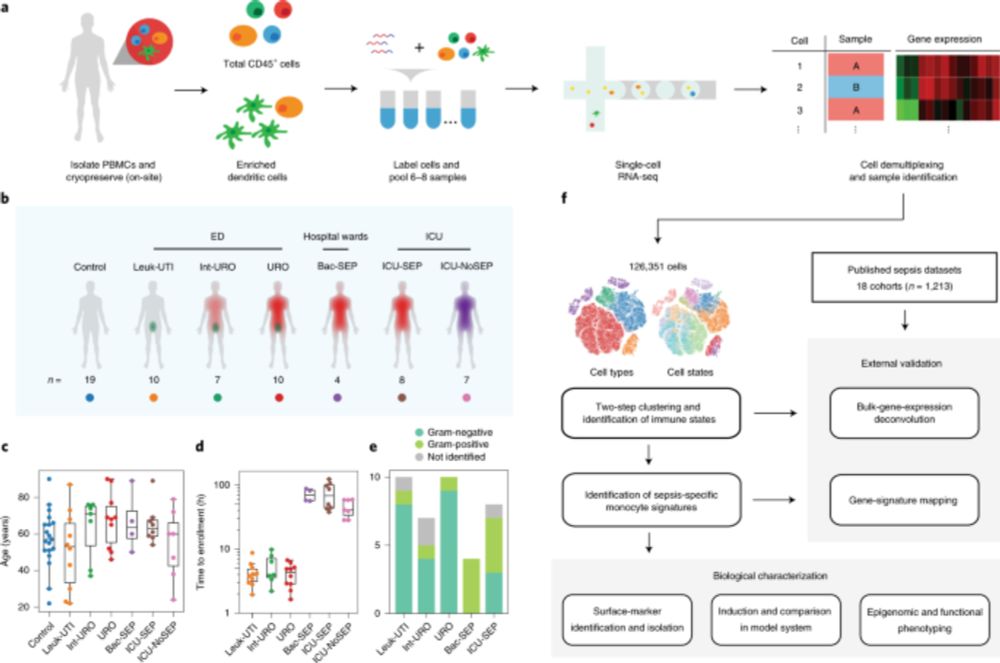
Single-cell transcriptomic analysis identifies a distinct gene signature associated with peripheral monocyte populations that distinguishes people with sepsis from those with sterile inflammation and ...
We piloted this in sepsis, to show a big advantage of the time-savings in ID: you can’t plan when a patient w sepsis will seek care, so to study it w scRNAseq, you need CRCs on call any time day or night, ready to do 2+ hrs of work for each pt. (We did this for COVID! But it’s hard.) Now? <15 min.
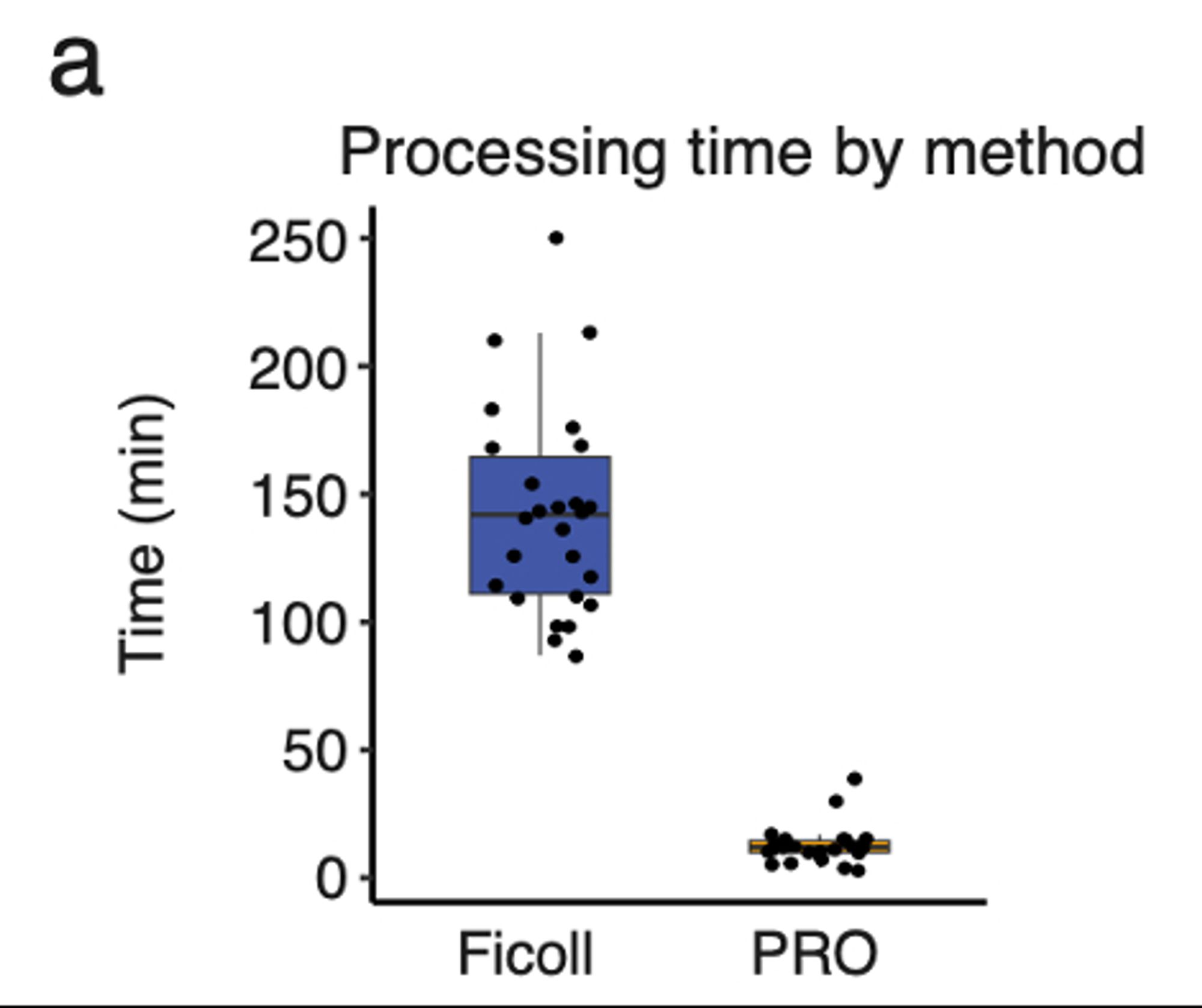
The post-thaw process involves magnetic depletion of RBCs, then FACS to isolate PBMCs (this step we normally do post-Ficoll too), then standard 10x LC & Seq. Standard QC metrics (% viable, genes/cell, UMIs/cell) look very similar to Ficoll, as do CITE-Seq metrics.
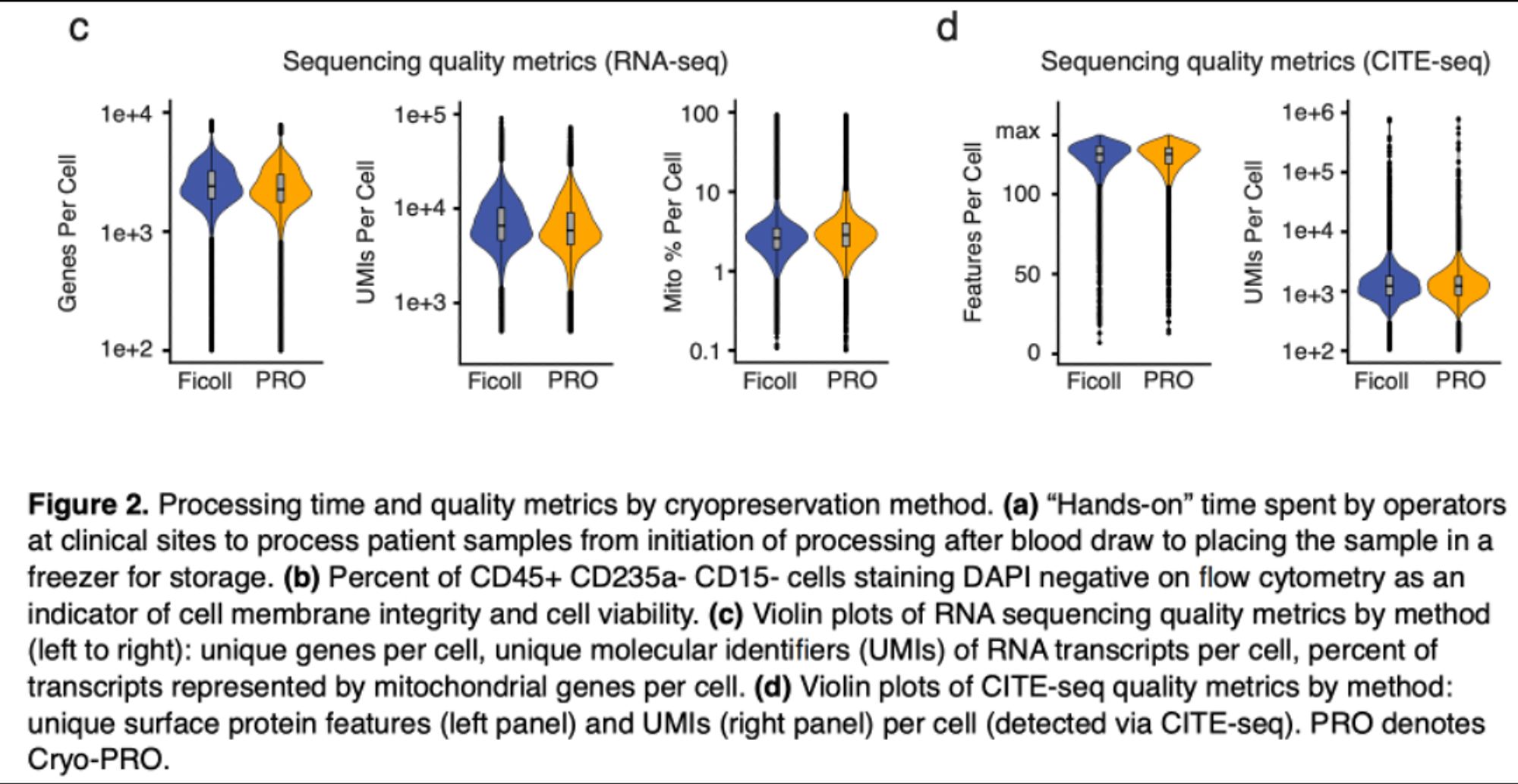
Turns out if you just add DMSO to 1-2 mL of whole blood & freeze immediately, it takes a few minutes (compared with >2 hrs for a Ficoll prep), and we worked out a post-thaw protocol that gets equivalent scRNA-seq data from immune cells (PBMCs), & can be run in batch at a centralized facility.
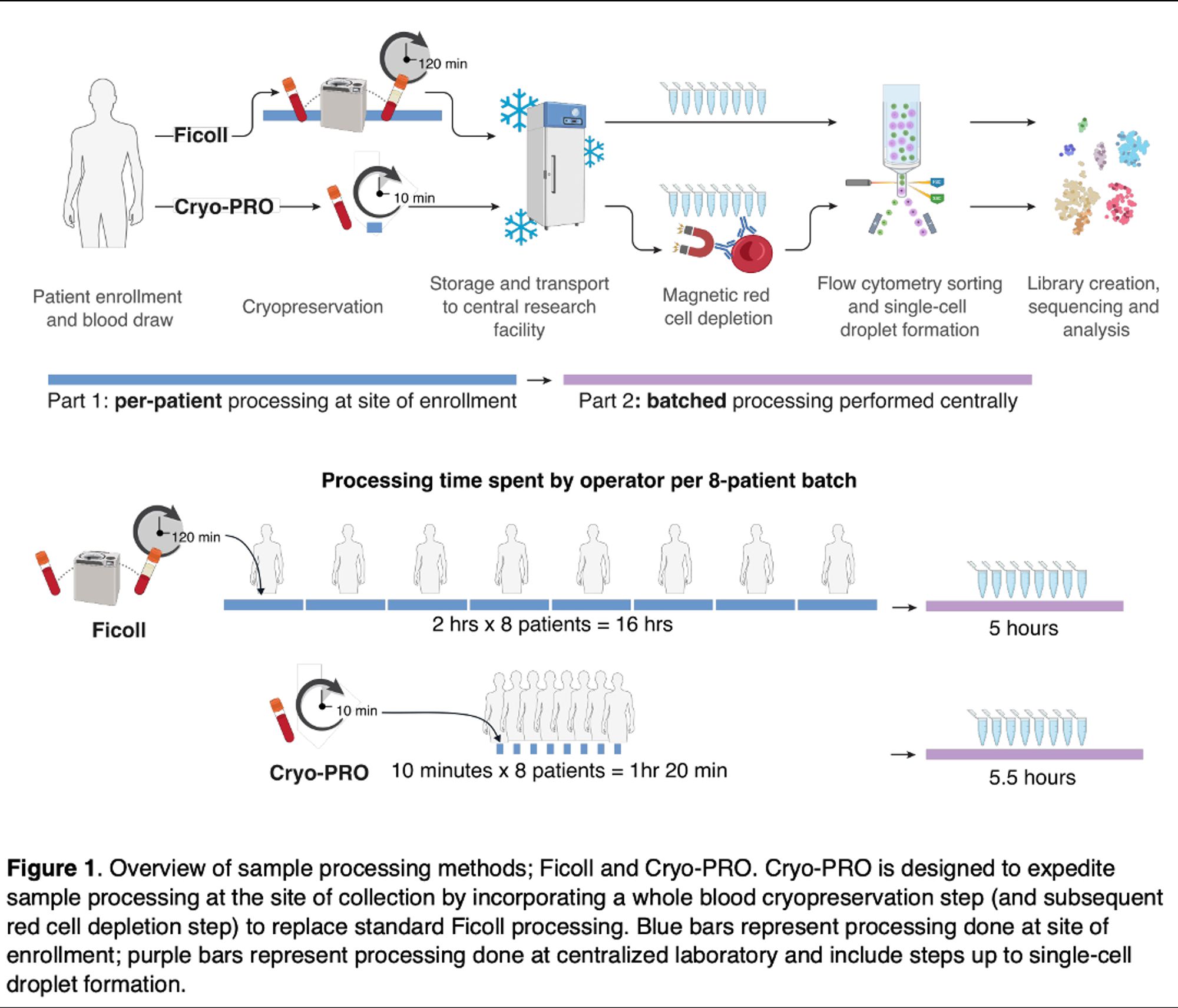
🧵 on a new preprint from my lab! scRNAseq has changed the way we view heterogeneity in the immune system. But in clinical settings, it’s really hard to do: processing blood for scRNAseq takes 2 hrs + expertise & equipment that clinical research staff often lack. We found an easier way!

medRxiv - The Preprint Server for Health Sciences
They may not come up well in single cell rnaseq data because they’re full of RNAses that chew up transcript within the droplets… there’s been some success with mouse tissues seeing eos on scRNAseq but very little in human and it takes lots of supplemental RNAse inhibitors.
Unlocking disease associations during prefrontal cortex development with scRNAseq #singlecell https://www.researchsquare.com/article/rs-4948061/latest