Very nice article by Danielle Gerhard in The Scientist on “The Dynamic Lives of Intrinsically Disordered Proteins” with quotes from Gabi Heller, @alexholehouse.bsky.socialwww.the-scientist.com/the-dynamic-...
Shapeshifting proteins challenge a long-standing maxim in biology.
Also, check out the complementary preprint from @lindorfflarsen.bsky.socialwww.biorxiv.org/content/10.1...

bioRxiv - the preprint server for biology, operated by Cold Spring Harbor Laboratory, a research and educational institution
This opens the door to the design of precise IDR-associated mutations that may fall outside of canonical binding motifs but will still enhance/suppress intermolecular interaction.
Finally, we see this approach as complementary to existing computational methods (structure prediction, simulations, sequence feature analysis, etc.) rather than replacing any of those. It fills a gap we need, and we hope it will be useful to others.
There are lots of caveats and limitations associated with this, and we encourage folks to consider those, but we also think this offers a way to build hypotheses and explain extant data. Predictions can be made over at finches-online.com
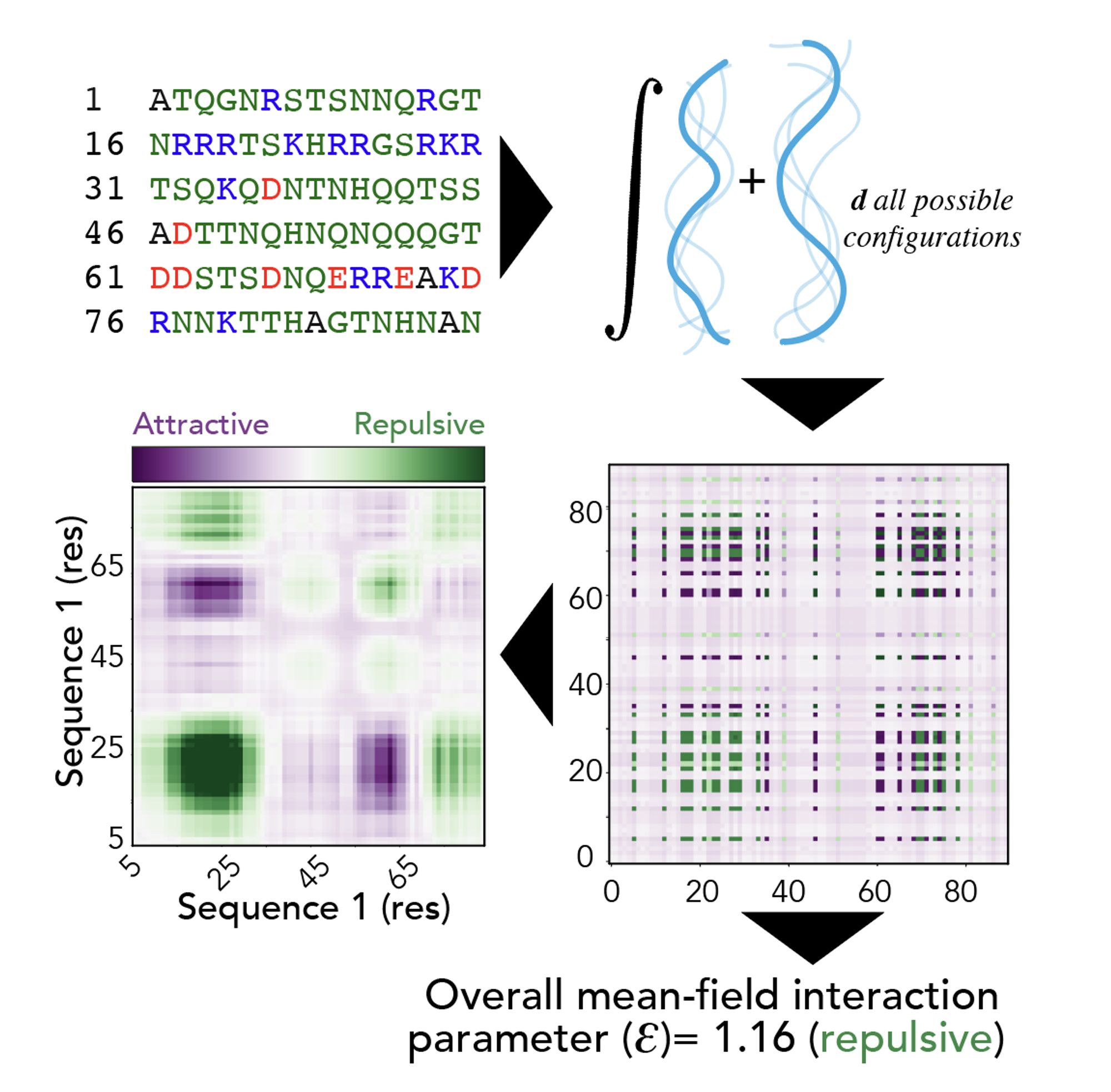
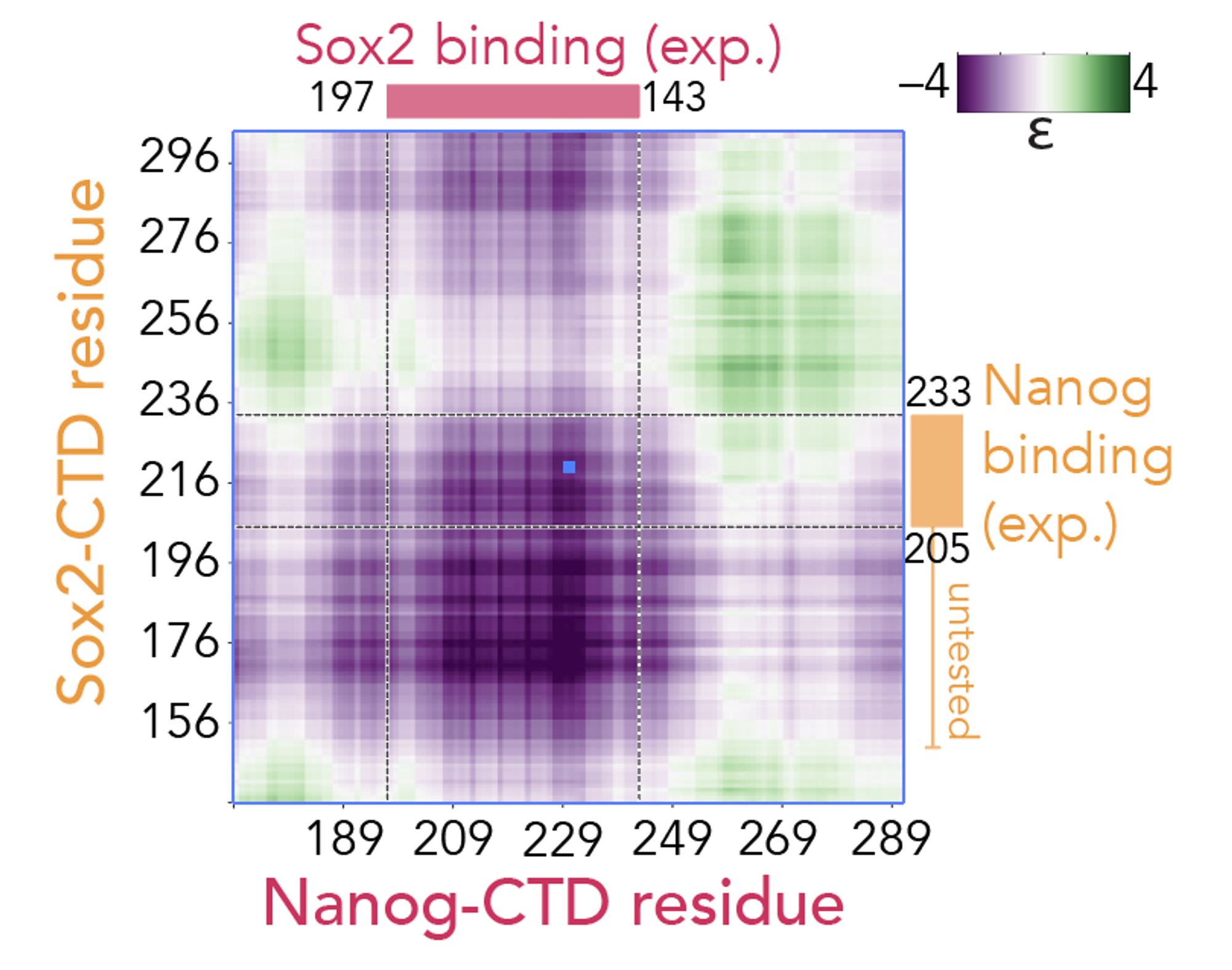
Much more exciting to me is our ability to predict heterotypic interaction between two IDRs (or an IDR and a folded domain surface) with residue resolution, making it simple to build testable hypotheses as to how an IDR may interact with a partner
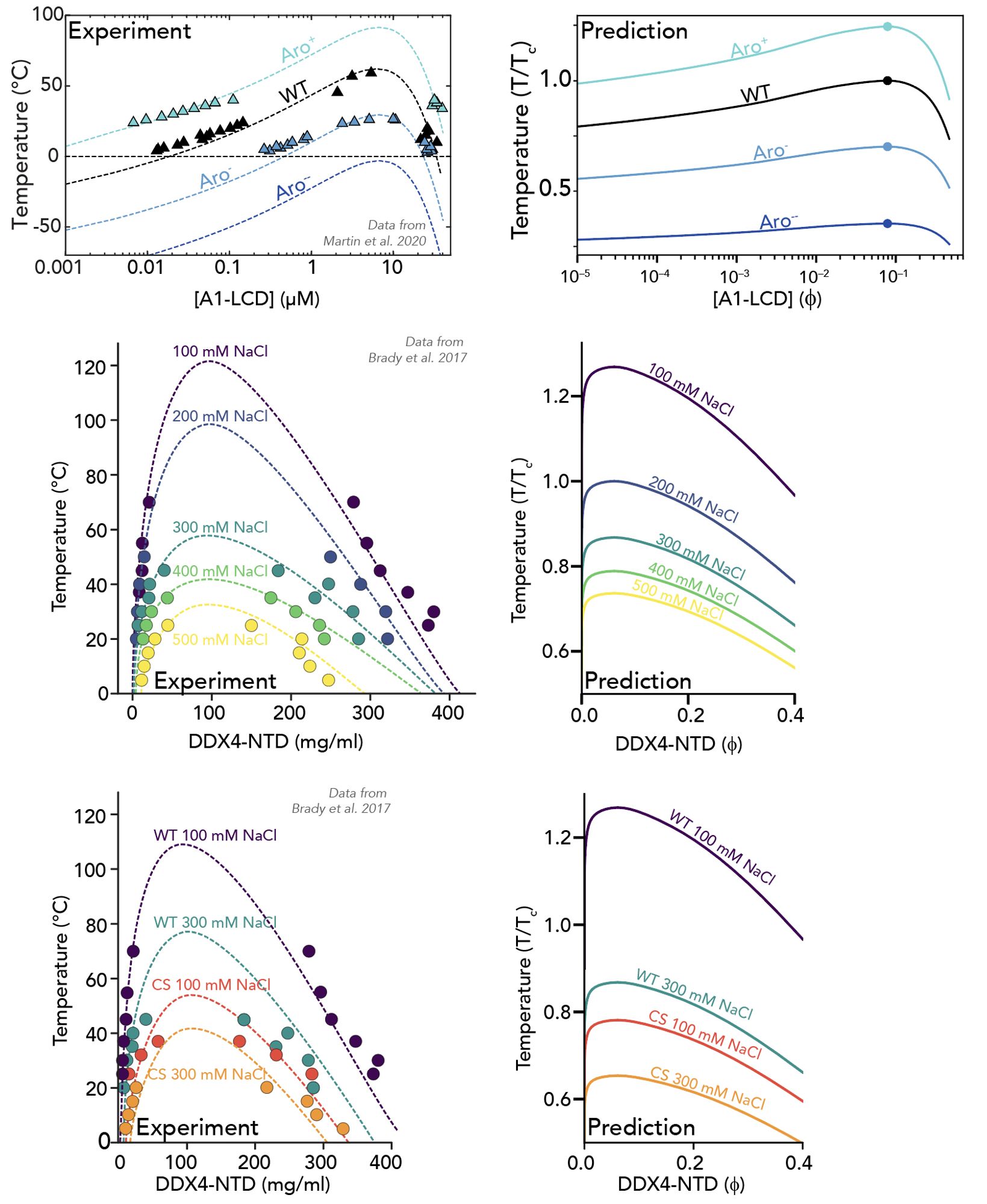
This approach allows us to do a few things. One of these is we can (qualitatively; lots of caveats) predict full phase diagrams from sequence
Excited to share new work from Garrett Ginell et al where we present an approach for predicting intermolecular interaction driven by disordered regions. We focus on how chemical complementarity can contribute to and even dictate intermolecular interactions www.biorxiv.org/content/10.1...