Lastly we are interested in cases where our predictions fail and unpublished data linking transmitters, neuropeptides, gap-junctions, post-synaptic receptors, co-transmisison combinations to EM. Please get in touch (Alexander_Bates[at]hms.harvard.edu) with any insights.
Thank you Lou Scheffer, Clare Pilgrim, @vincentcroset.bsky.social@hhmi.bsky.social.
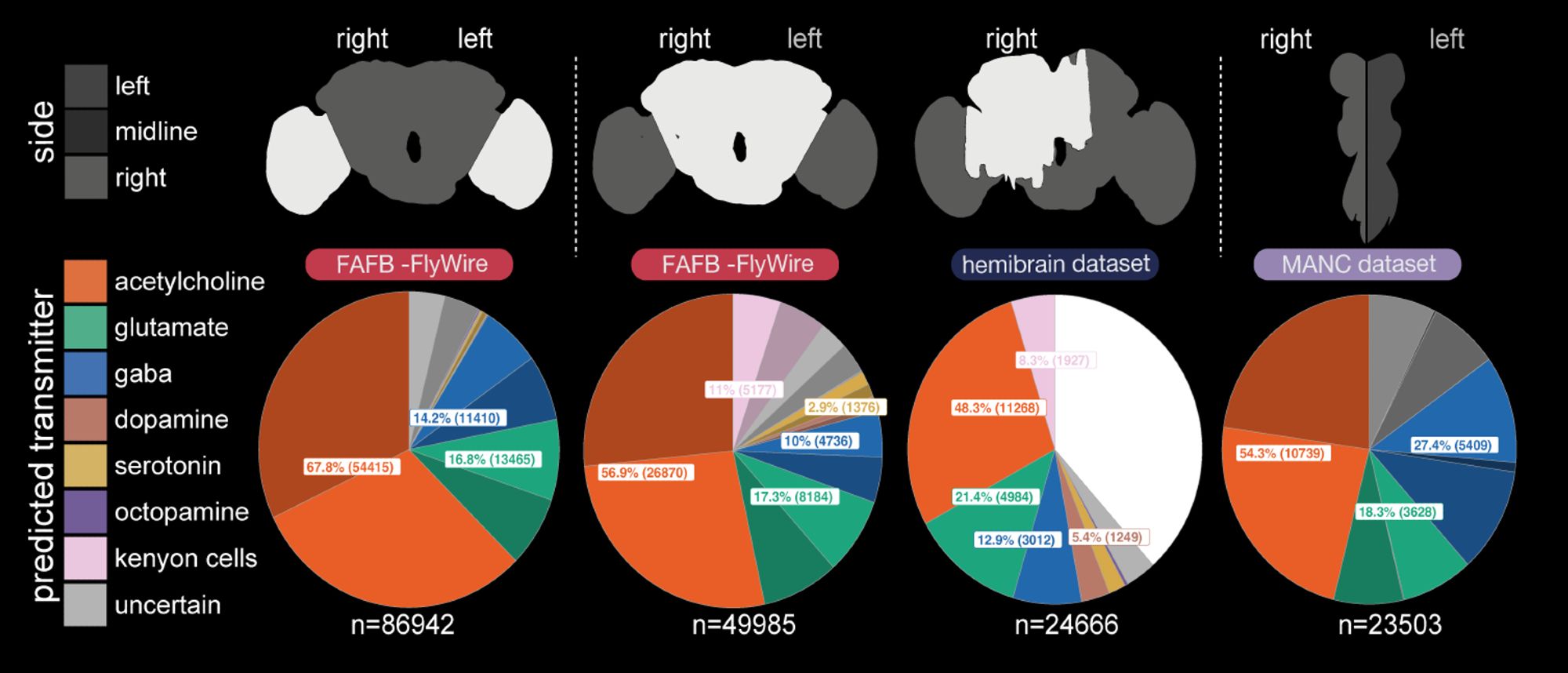
GT built from the FAFB (@dddavi.bsky.social@perlman.bsky.socialVirtualFlyBrain.org.
Welcome to Virtual Fly Brain (VFB) - an interactive tool for neurobiologists to explore the detailed neuroanatomy, neuron connectivity and gene expression of Drosophila melanogaster. Our goal is to ma...
Nils Eckstein, Andrew Champion, Michelle Du and Jan Funke for the ML, biology w/ Yijie Yin, Philipp Schlegel, Kathi Eichler, @martamcosta2, @gsxej, Volker Hartenstein, data w/ Sven Dorkenwald, Arie Matsilah, Claire McKellar, Amy Sterling, Mala Murthy, Sebastian Seung, the Janelia Annotation Team.
But we feel our work is a big step forward. Our data (zenodo.org/records/1059...github.com/funkelab/syn...codex.flywire.aiVirtualFlyBrain.org.
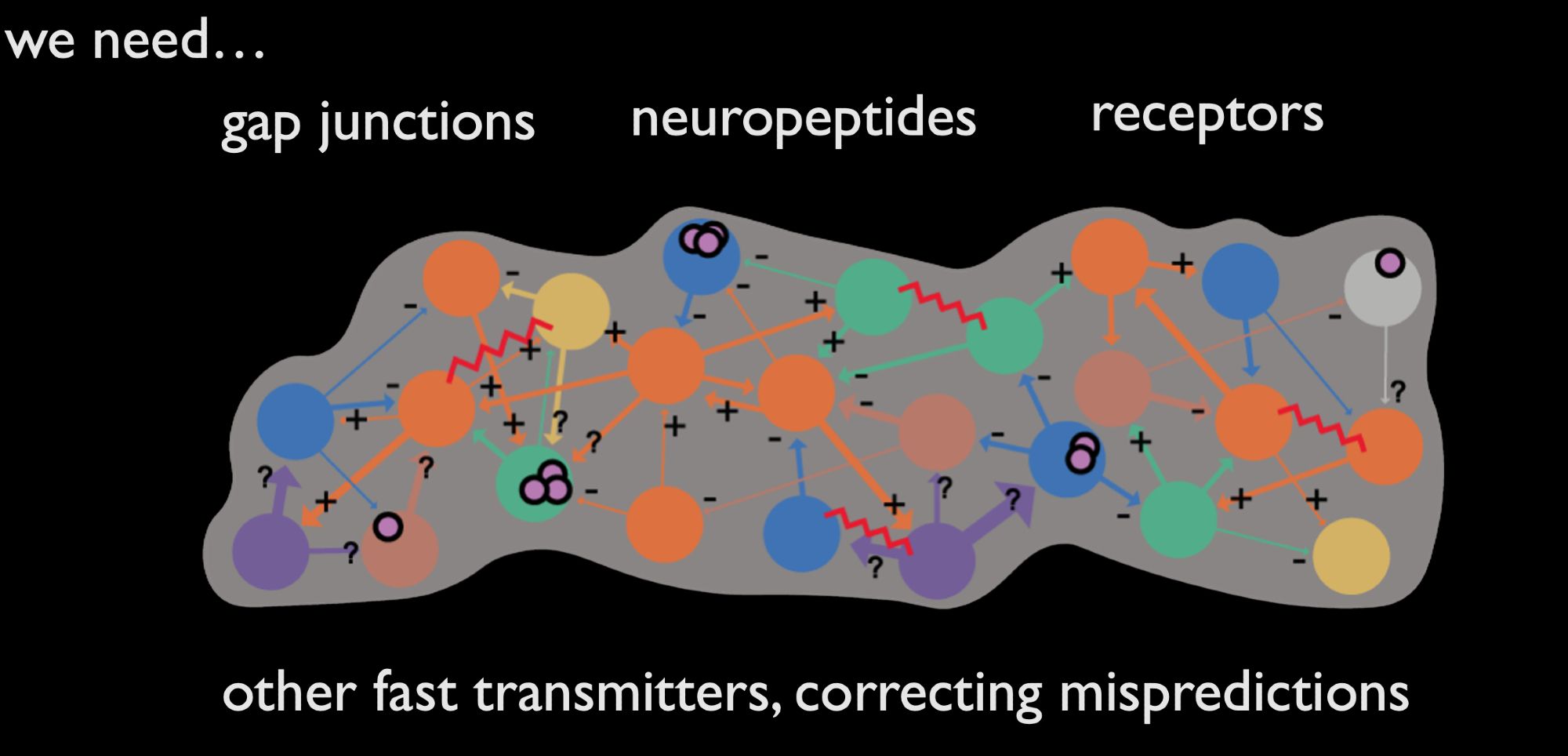
Not quite mission complete. We want more annotations. E.g. receptor localisation delineates excitatory from inhibitory glut connections, moving our +/ - labels to individual connections. Their annotation could enable ML to extract this molecular information from EM at scale.
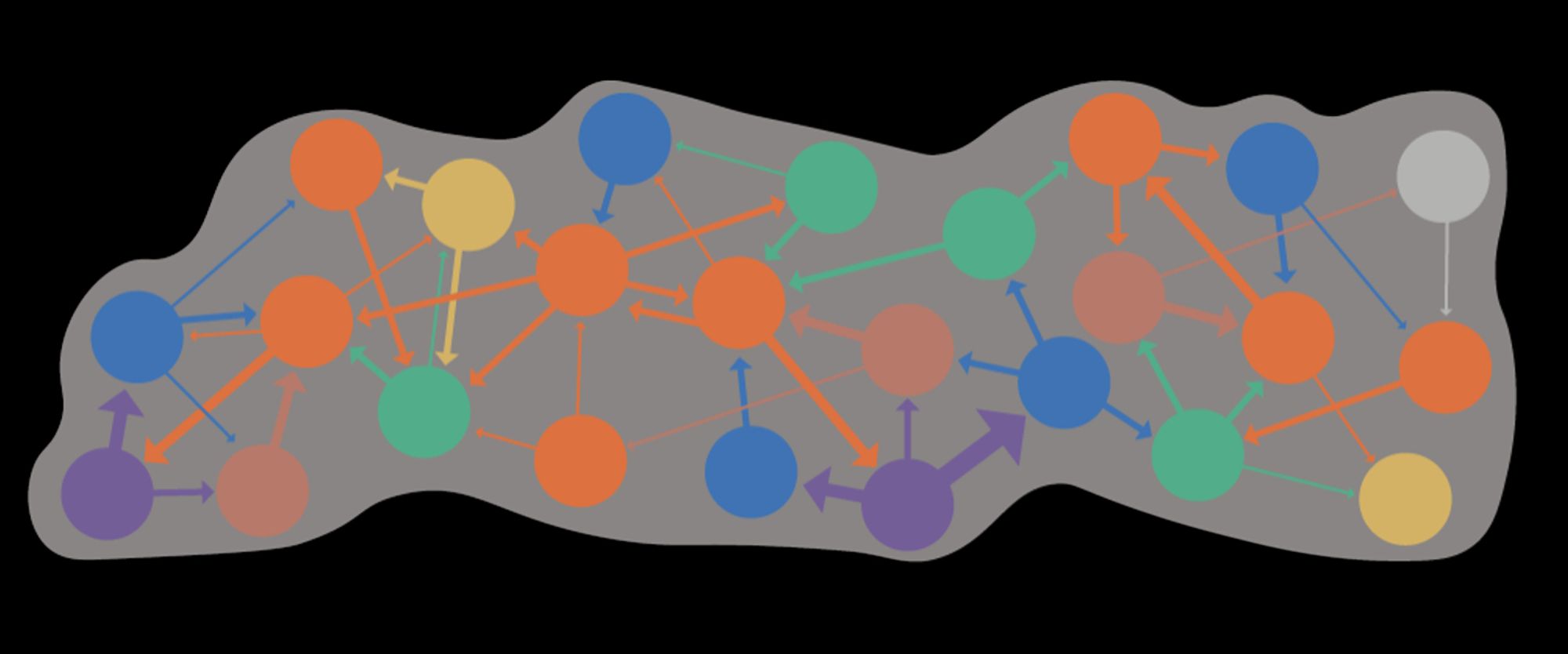
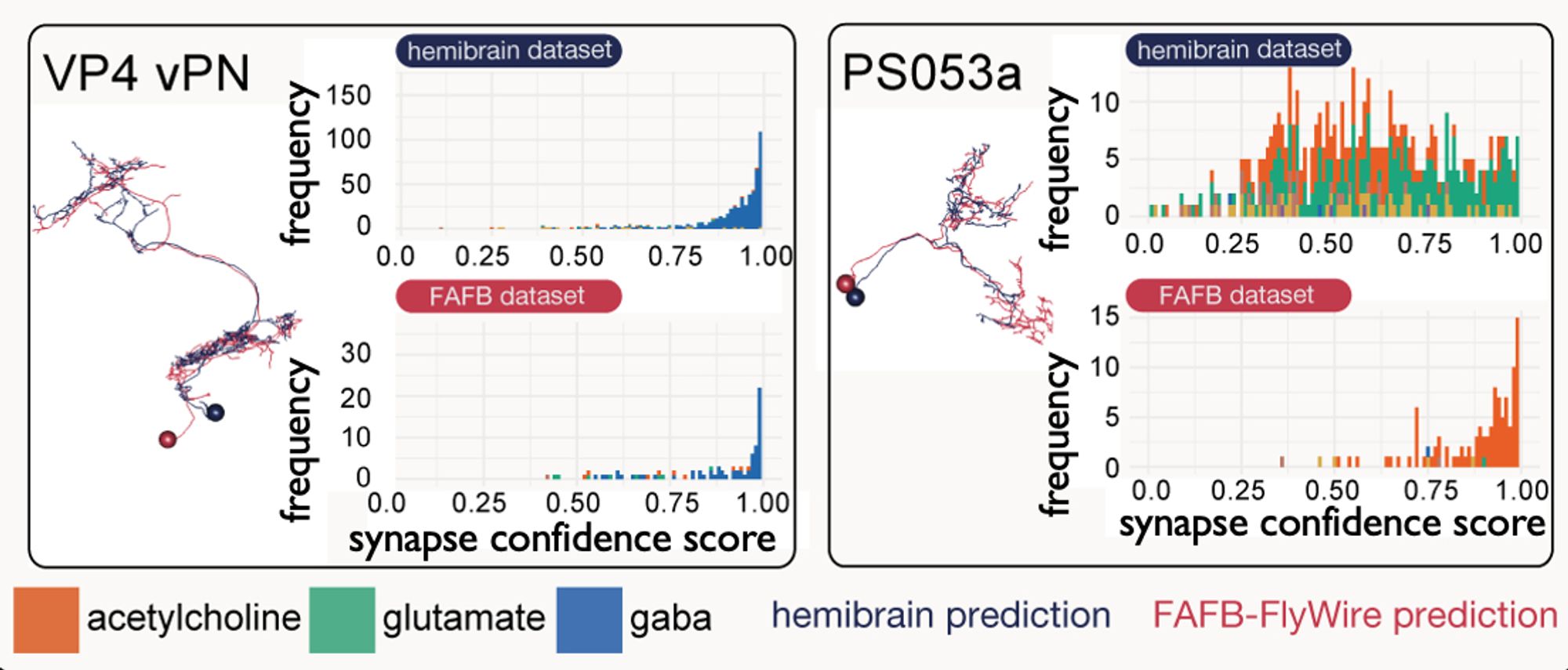
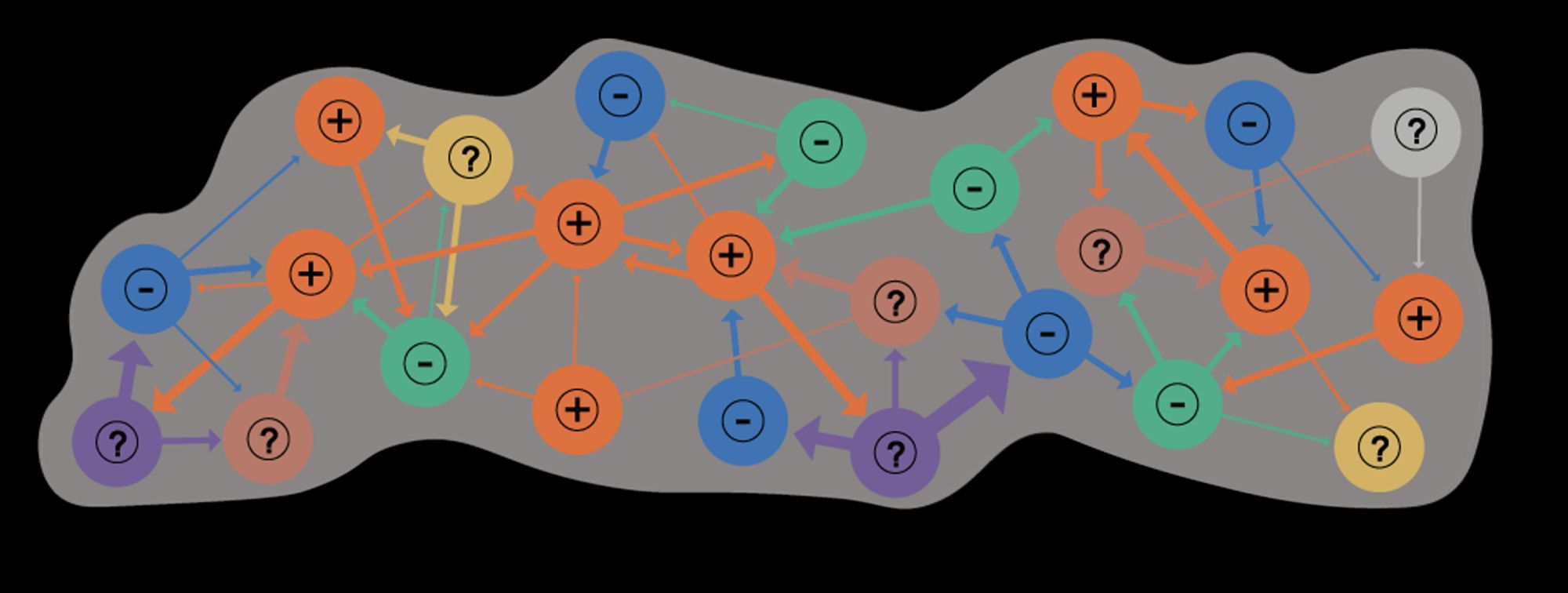
Which proxy for synaptic signs!... With major assumptions, e.g. no co-transmission, and that glutamate is inhibitory - but, it seems ~80% of brain neurons may express both excitatory and inhibitory GluRs, and co-transmission esp. in monoaminergic neurons is reasonably common.
Yet overall, we had predictions which we felt were pretty good, across several data sets that together make up a whole (amalgam) Drosophila melanogaster nervous system! The neurotransmitter proportions turned out similar to those suggested by single-cell RNA-seq studies.