This work wouldn't be possible without our fantastic collaborators @katholt.bsky.social Drs Sylvain Brisse, Jon Monk and Martin Rethoret-Pasty. Plus important contributions from the wonderful Helena Cooper! 🙏
This is cool 😎 but why should you care? Our data shows that Kp clones have different metabolic capabilities (and we are probably underestimating this) This could drive differences in virulence and fitness e.g. clone-specific traits have reported roles in gut competition!
We propose that the metabolic diversity between clones facilitates their ongoing co-existence in the population by 1. reducing nutrient competition 2. promoting commensal interactions consistent with a model of negative-frequency dependent selection
When we tested some of these clone-specific core substrates in the lab, we found evidence of reciprocal cross-feeding between clones!
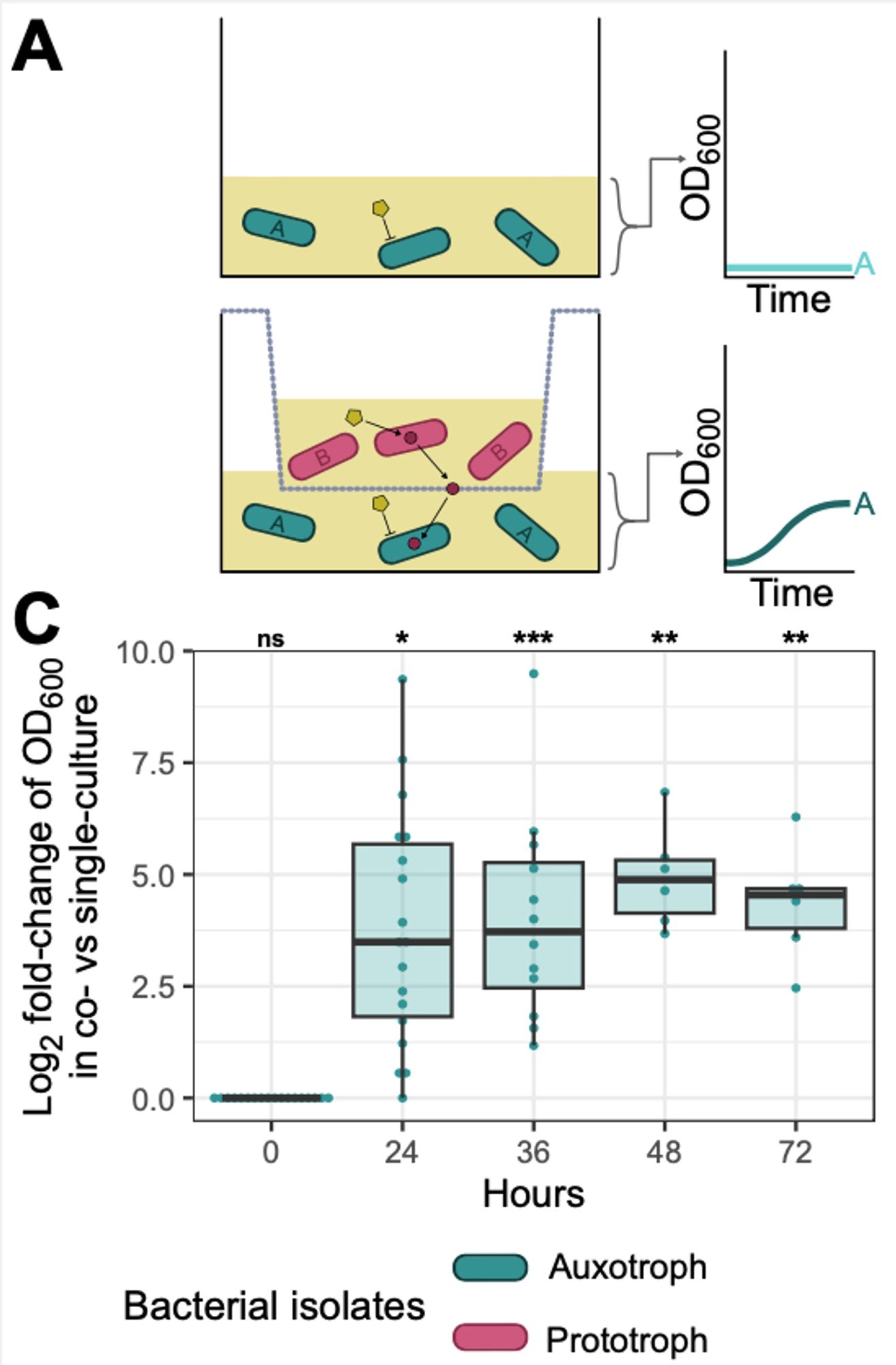
When we took a close look at 48 common clones (sub-lineages) we found each had a unique predicted phenotype profile. We defined clone-specific core phenotypes- some common to many clones, some rarer e.g. global MDR SLs 258 and 307 could use fructoselysine, but SL147 couldn't
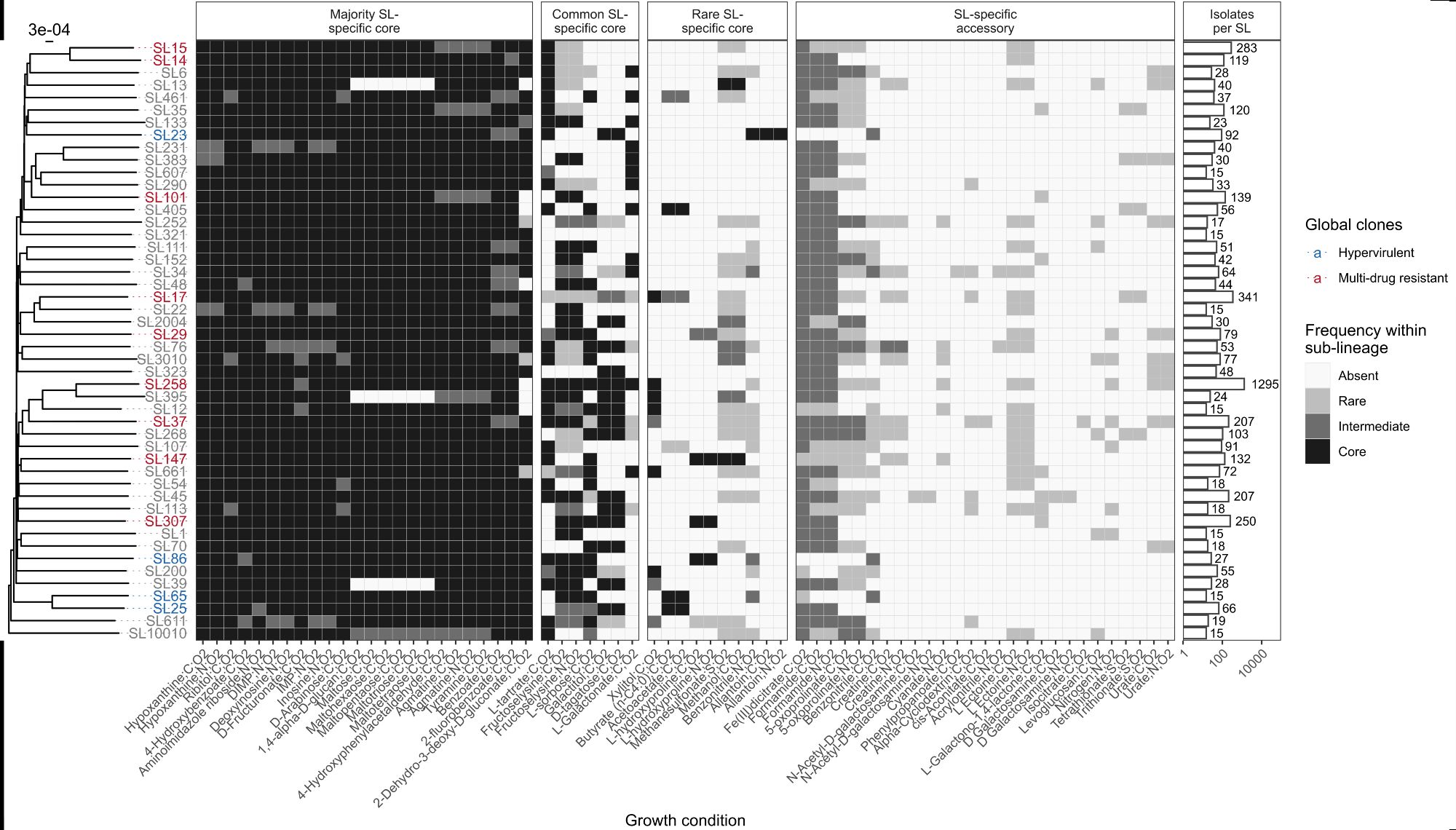
But there was variation within species too! This was structured so that more closely related isolates shared more similar sets of metabolic traits.
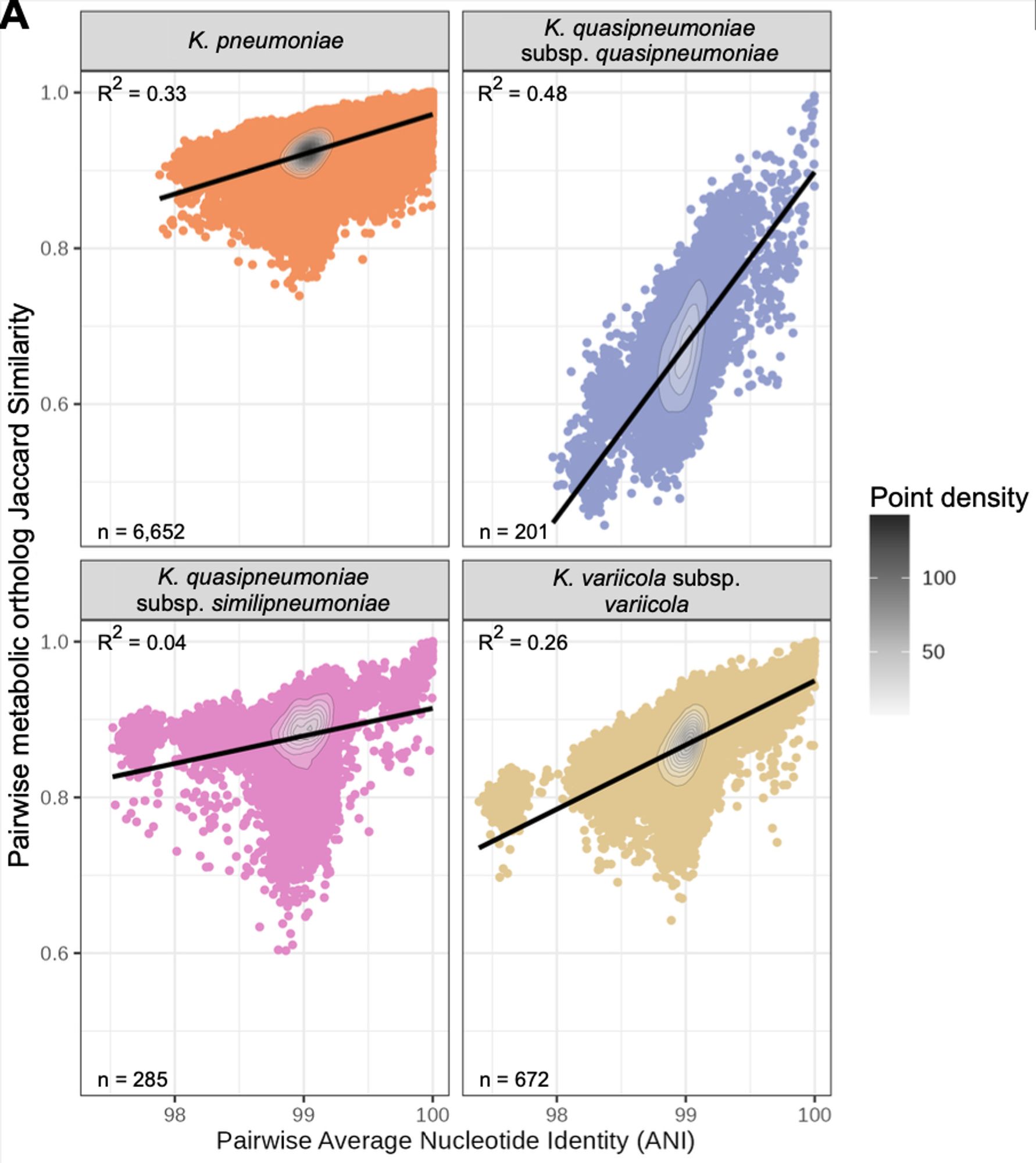
We simulated 1300 growth phenotypes for each isolate (different carbon, nitrogen, phosphorus, sulfur sources in minimal media +/- O2). Phenotype profiles differed by species - as we expected - and were generally consistent with formal species definitions.
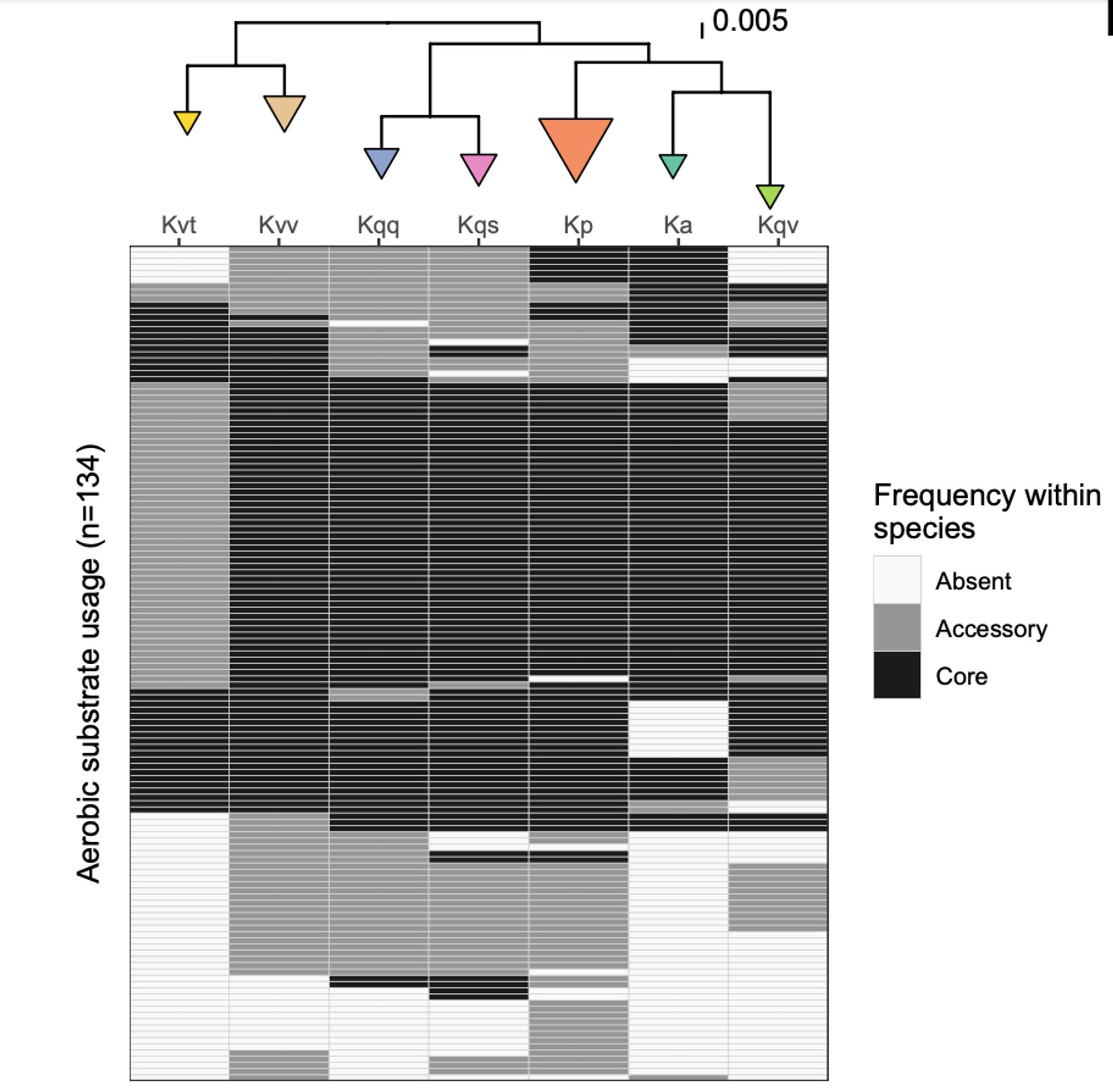
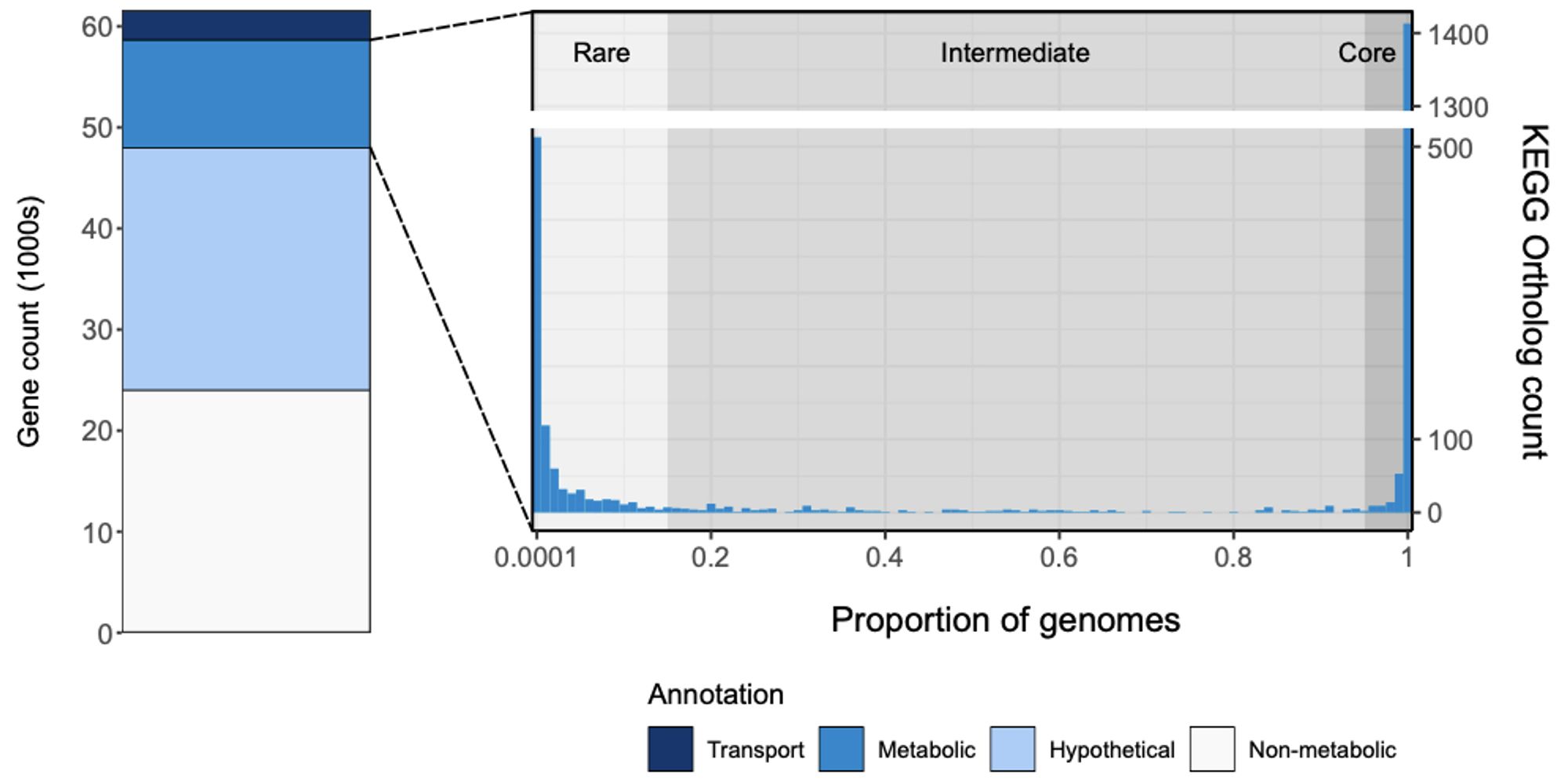
We used a combination of genomics and metabolic modelling of >7K Klebs pneumo complex isolates to explore diversity and distribution of metabolic traits. Most individual traits are either core or very rare - just like genes in the total pan-genome.
Ever wondered why there are so many co-circulating #klebsiella@bananabenana.bsky.socialdoi.org/10.1101/2024...

bioRxiv - the preprint server for biology, operated by Cold Spring Harbor Laboratory, a research and educational institution