correction, conda is open source, miniconda is free but not OS. Only the repositories managed by Anaconda fall under this licensing scheme: docs.anaconda.com/miniconda/
From what I have read, as long as you remove the "defaults" channel then you are good to go without needing a license. Conda and miniconda are open source.
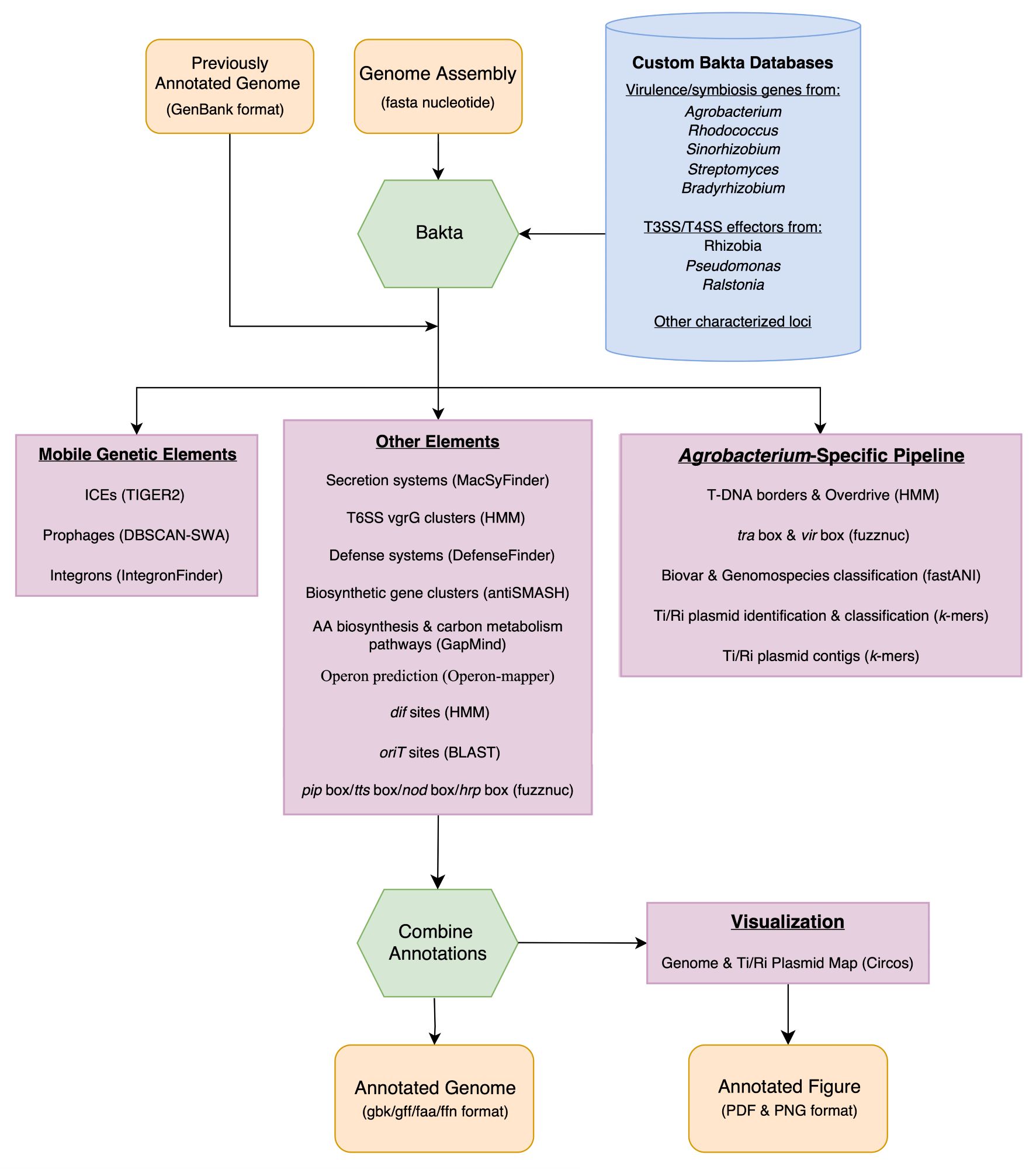
Our bacterial genome annotation tool Beav is now published in mSphere! The newest version also supports gbk files as input to add to previous annotations. Available on bioconda. journals.asm.org/doi/10.1128/...
The Dept. of Botany and Plant Pathology at Oregon State is hiring a TT Asst. Prof. in Nematology. Come be my colleague in beautiful Oregon! files.cqls.oregonstate.edu/Weisberg_Lab...
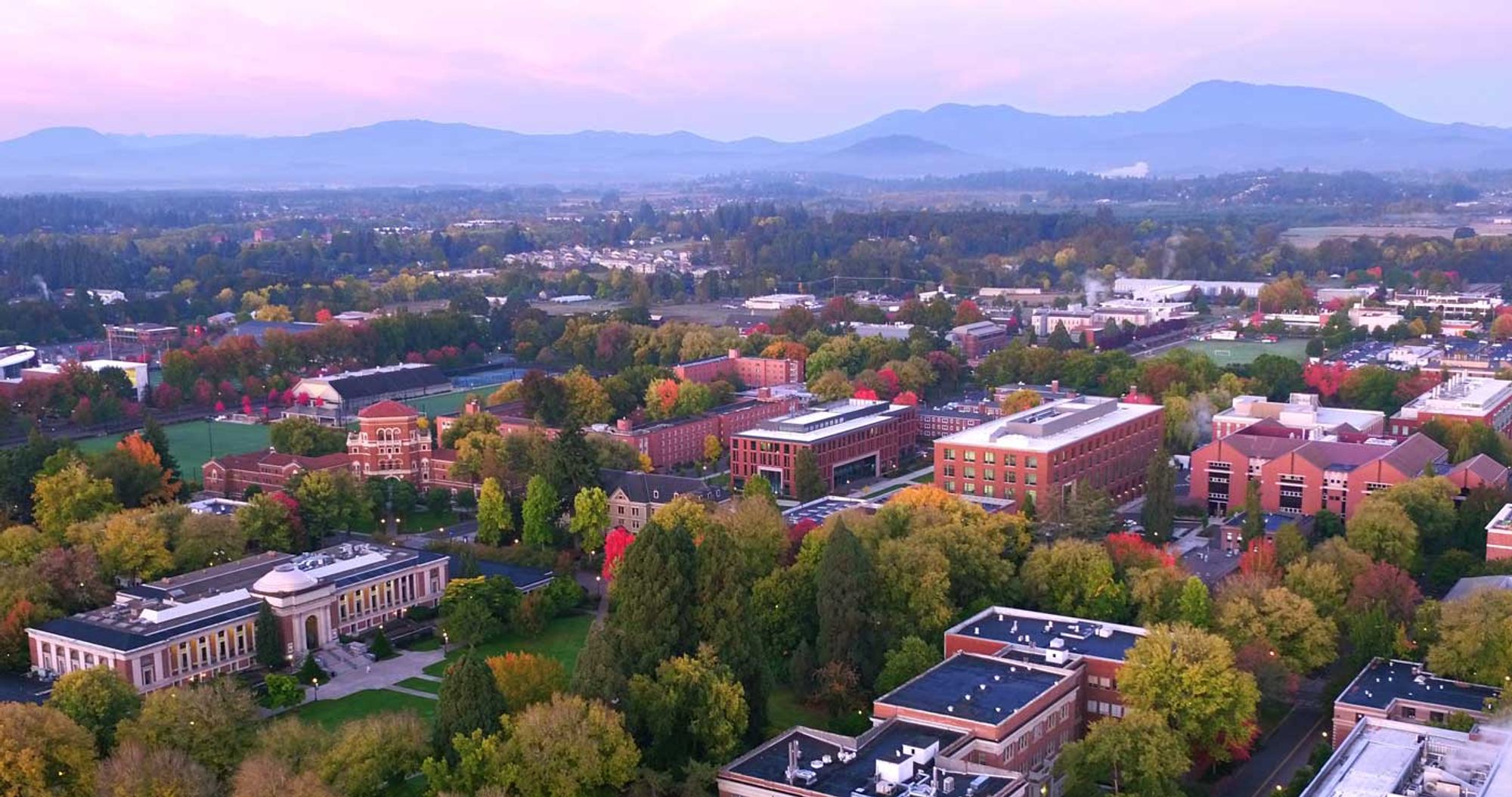
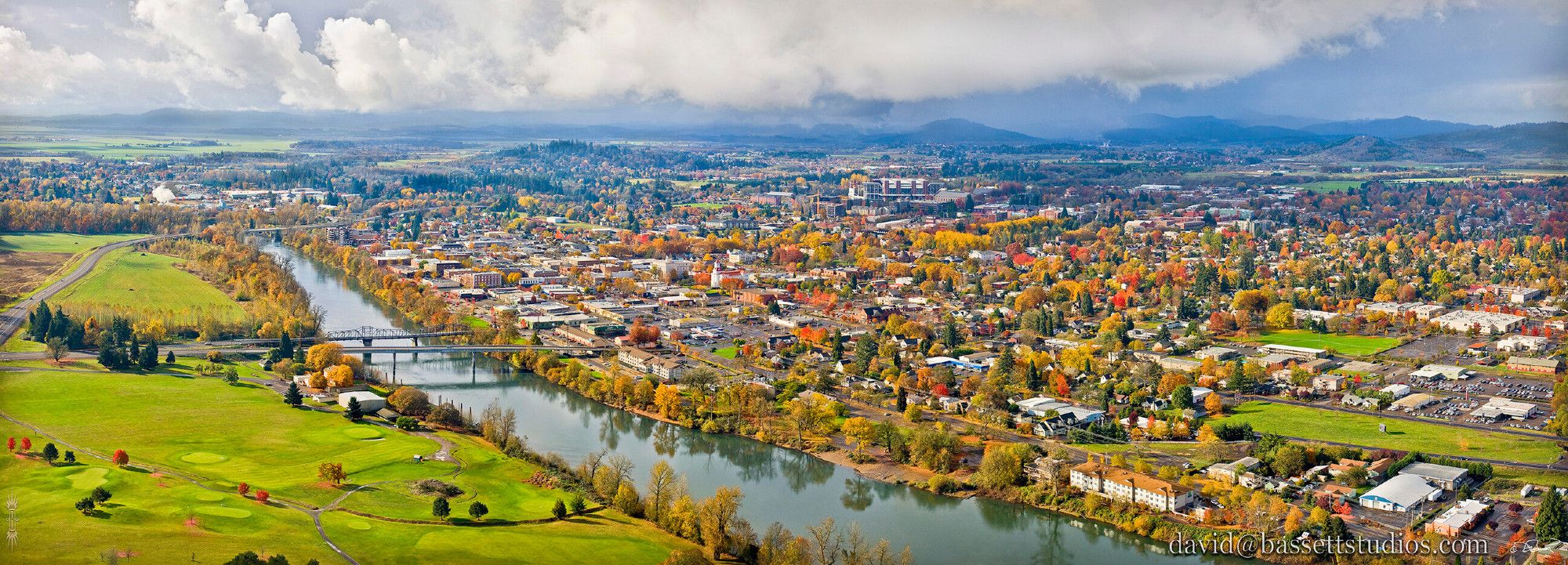
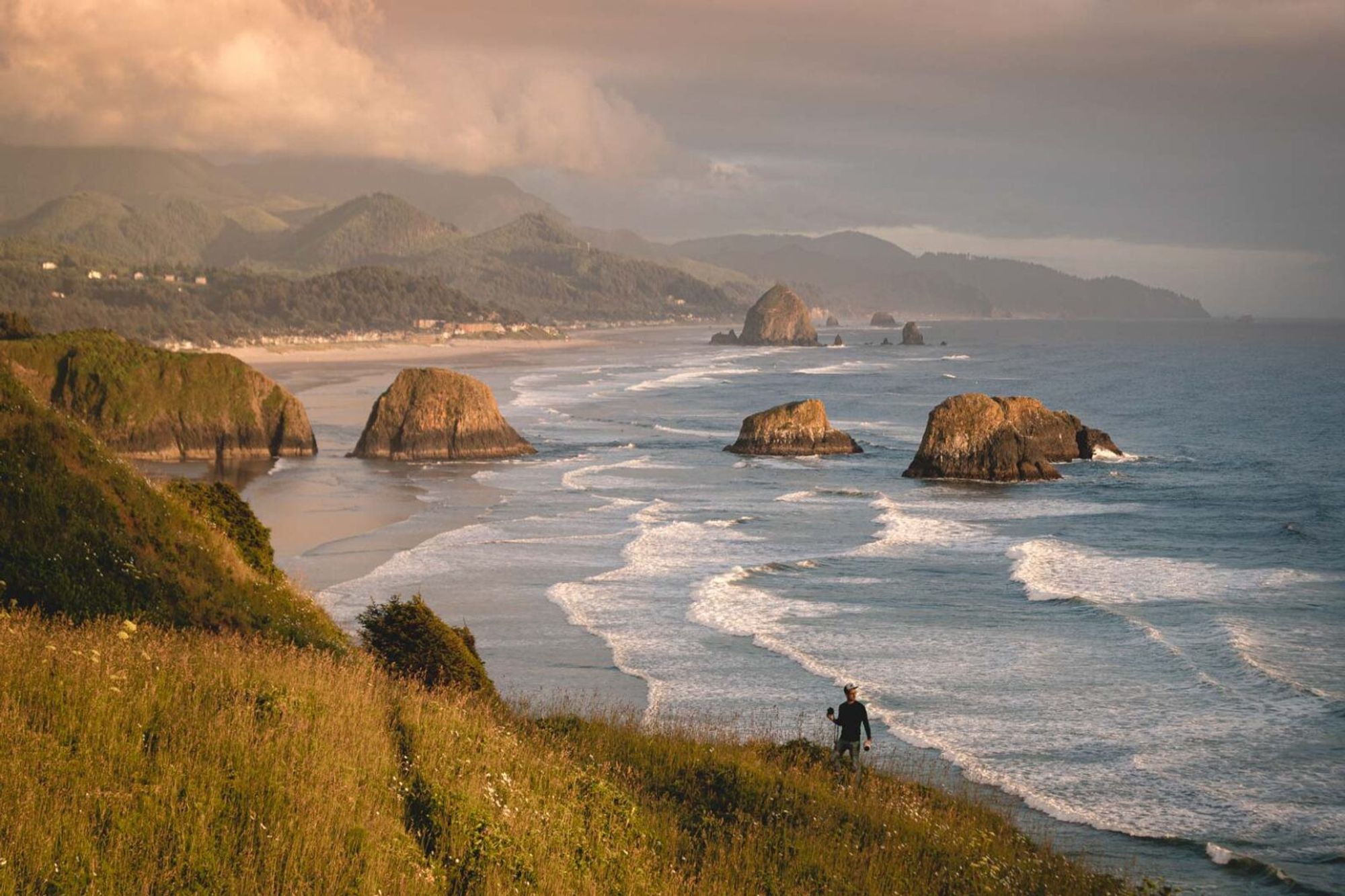




We’re trying to make the G3 journal a home for all your fun bacterial genetics stories! Please hit us up if you have a relevant manuscript to submit!

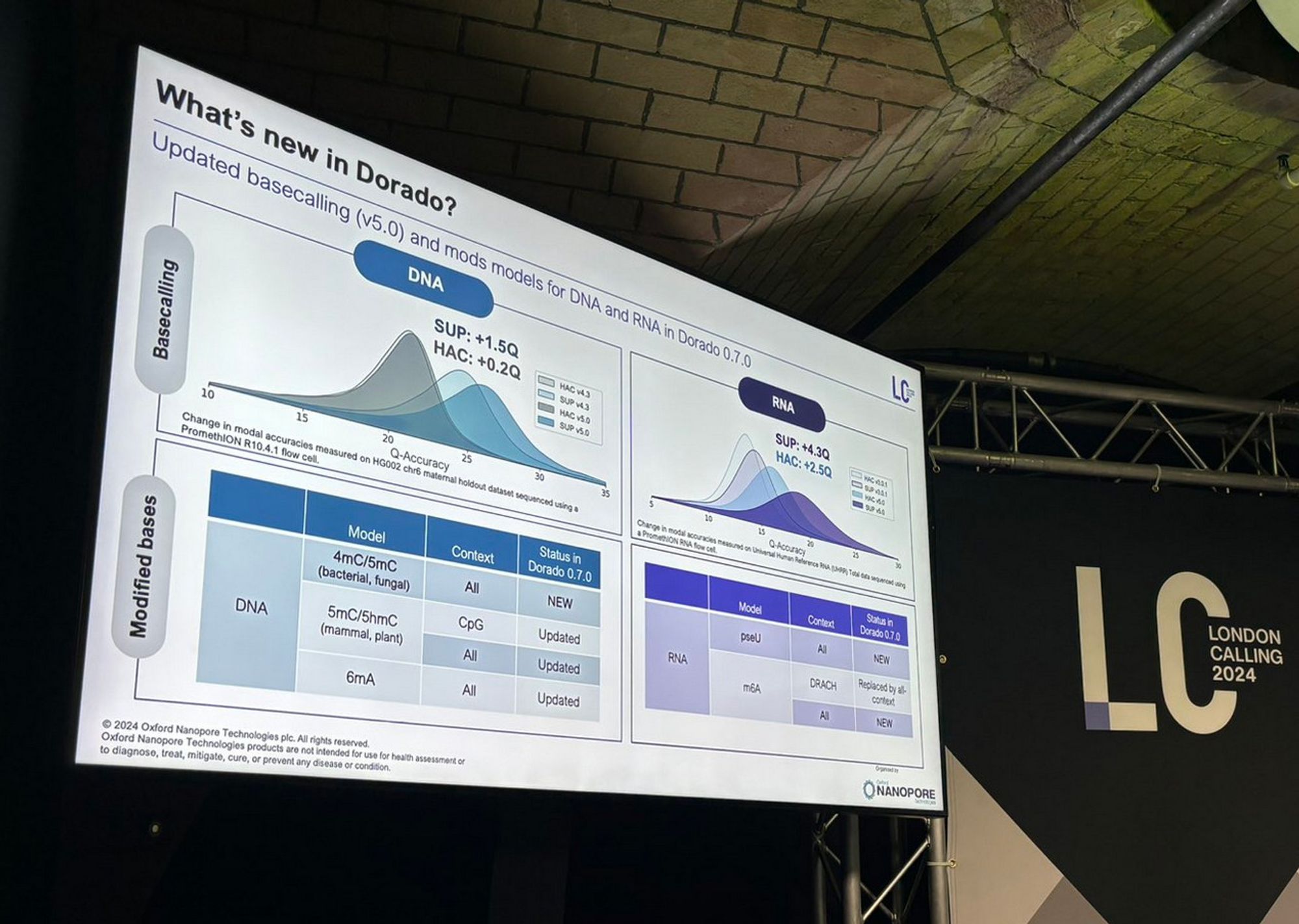
It's the most wonderful time of the year: the time when we all learn about the latest nanopore updates via screenshots of tweets of photos of slides Looks not disappointing, no sign of the accuracy wall! And dorado 0.7 is now out with the new models: github.com/nanoporetech...
Our new paper on siderophore gene expression heterogeneity across space and time in #Pseudomonas#CellReportswww.cell.com/cell-reports...
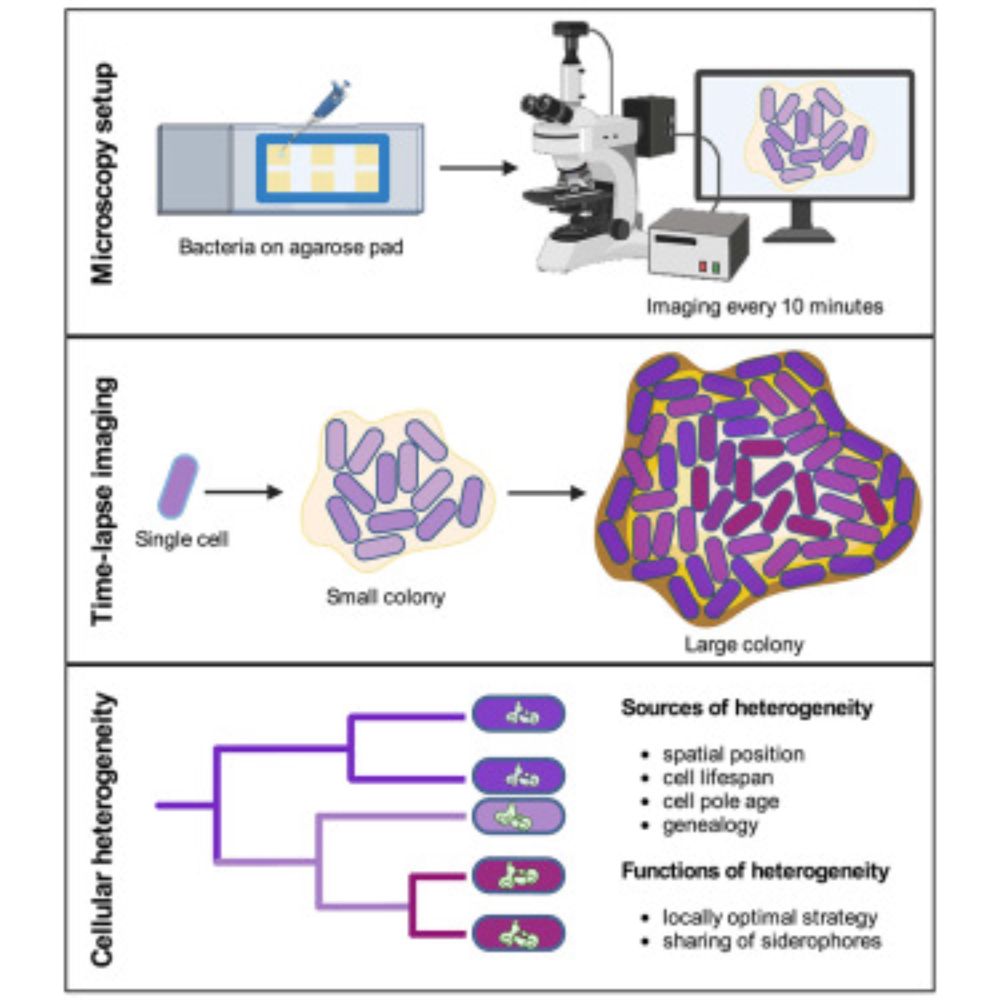
Mridha et al. show that the spatial positioning of cells within colonies and genealogical effects drive cellular siderophore gene expression heterogeneity in Pseudomonas aeruginosa bacteria. Functiona...
Something to keep in your quote armory: "...many papers in the literature report microbial associations that go against basic understanding of microbial ecology, some of which can be likened to reporting blue whales in the Himalayas or African elephants in Antarctica."
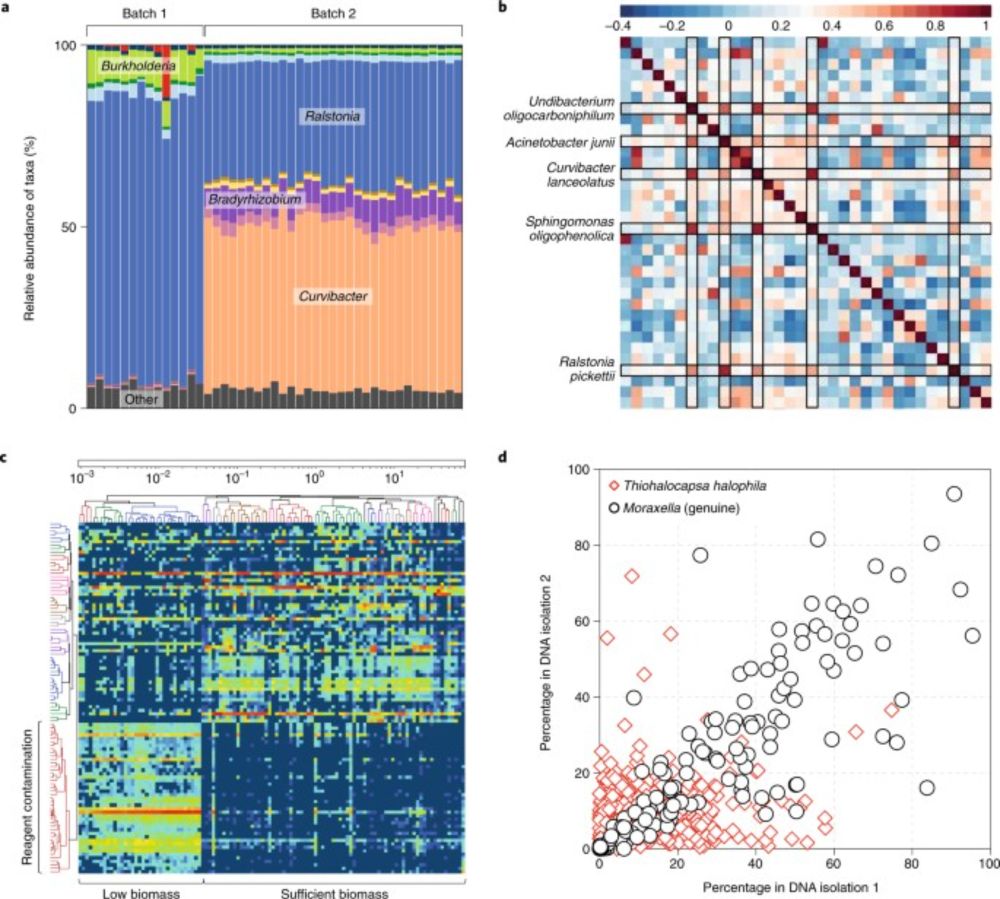
A noticeable part of the microbiome literature, especially that working with low-biomass samples, is plagued by reagent contamination. Here we describe visual, statistical, methodical and ecological techniques to facilitate recognition of signals that represent contamination.