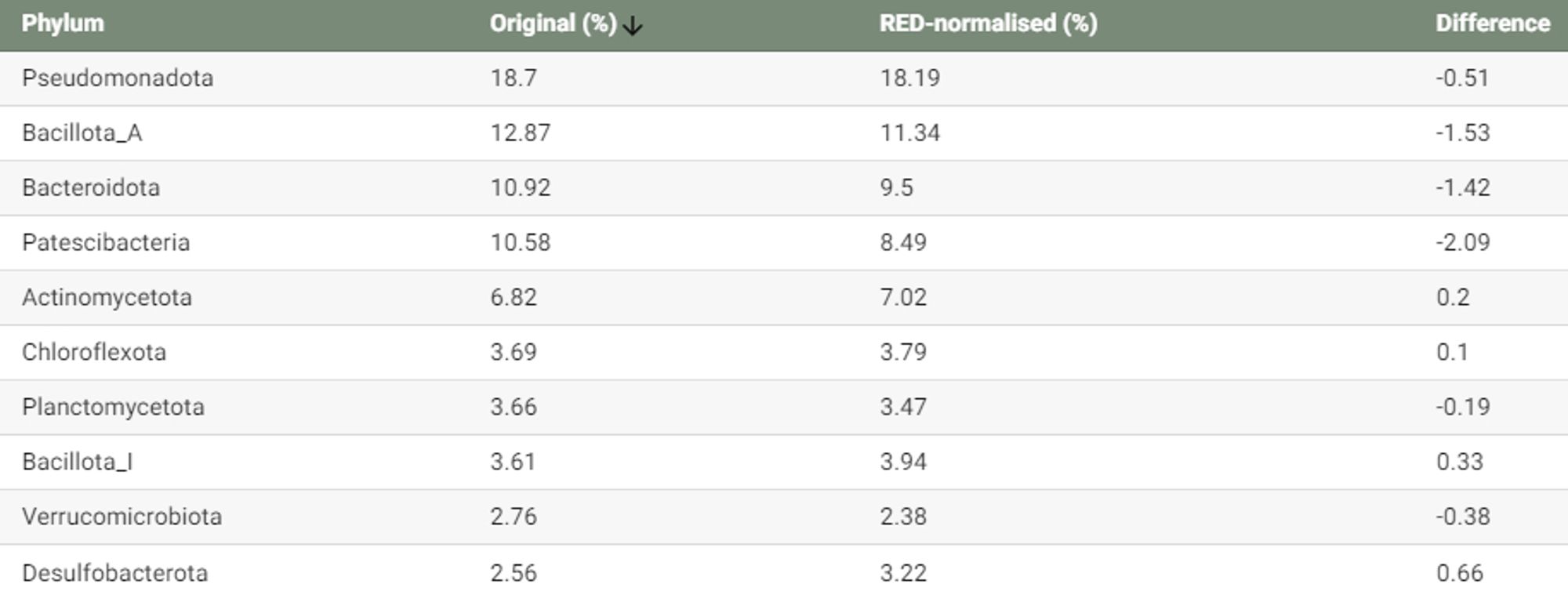
A single amplified genome catalog reveals the dynamics of mobilome and resistome in the human microbiome 17,202 SAGs in this work - mostly medium quality link.springer.com/article/10.1...
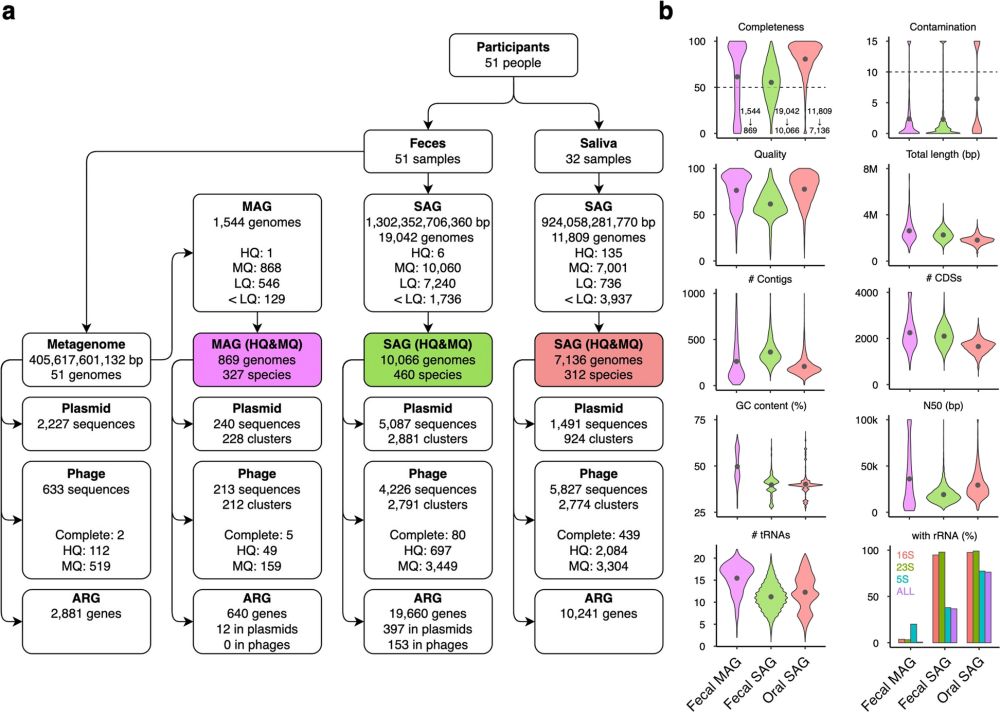
Background The increase in metagenome-assembled genomes (MAGs) has advanced our understanding of the functional characterization and taxonomic assignment within the human microbiome. However, MAGs, as...
Evolution of a bistable genetic system in fluctuating and nonfluctuating environments -in PNAS www.pnas.org/doi/10.1073/...
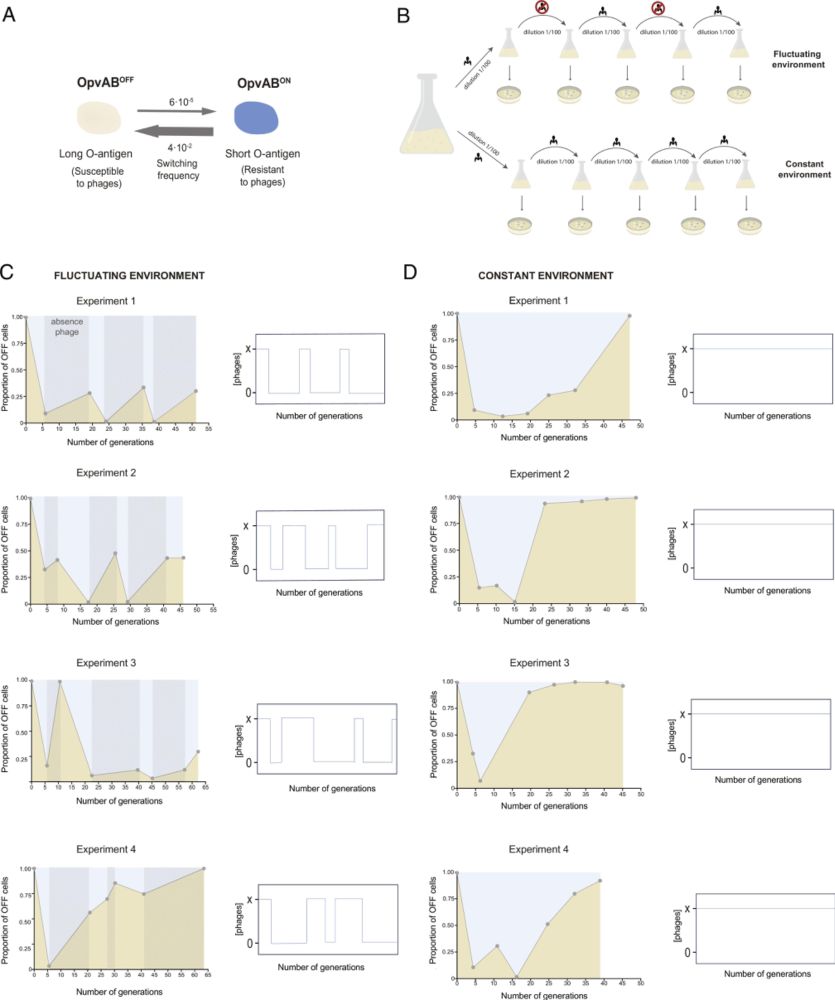
Epigenetic mechanisms can generate bacterial lineages capable of spontaneously switching between distinct phenotypes. Currently, mathematical model...
Early career researchers in microbial ecology, please consider joining the Early Career Scientist Reviewer Pool at ISME Journal. A great way to widen your scientific community. @ismeoffice.bsky.socialwww.isme-microbes.org/isme-early-c...
Are you going to #ISME19#anvioforms.gle/idoXJbN5bpUP... Thank you! 😇
The purpose of this form is to collect data on ISME19 posters that made use of anvi'o. Just so we, as in anvi'o users and developers, can recognize the kinds of applications people have used anvi'o fo...
microbiomejournal.biomedcentral.com/articles/10....
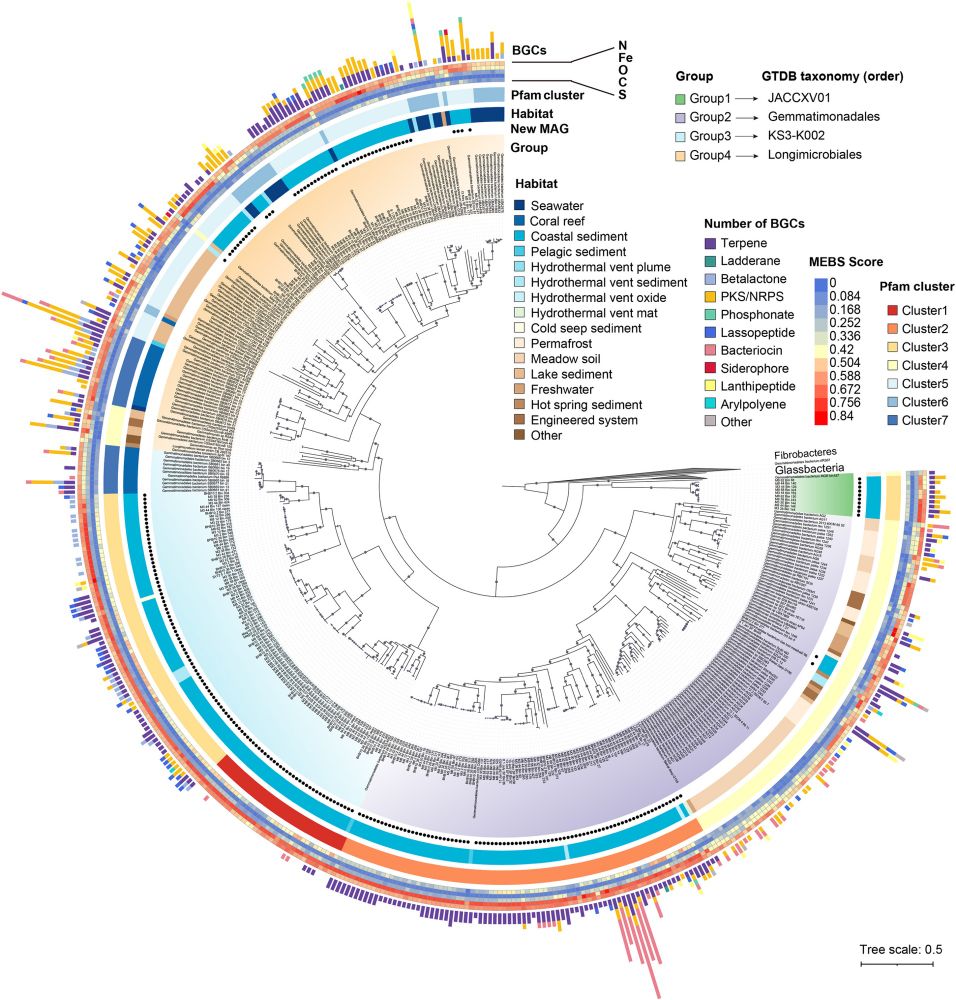
Background Gemmatimonadota bacteria are widely distributed in nature, but their metabolic potential and ecological roles in marine environments are poorly understood. Results Here, we obtained 495 met...
Wow! openalex.org looks like an amazing open alternative to Google Scholar. Citation counts etc. look very similar to GScholar on a quick vanity search. Check it out. Opens up a lot of possibilities for metascience.
Eight years later, I revisit the tree of life. “Future trees may need to distinguish not only between cultured and uncultured lineages, but between extant and extinct” rdcu.be/dPNpB
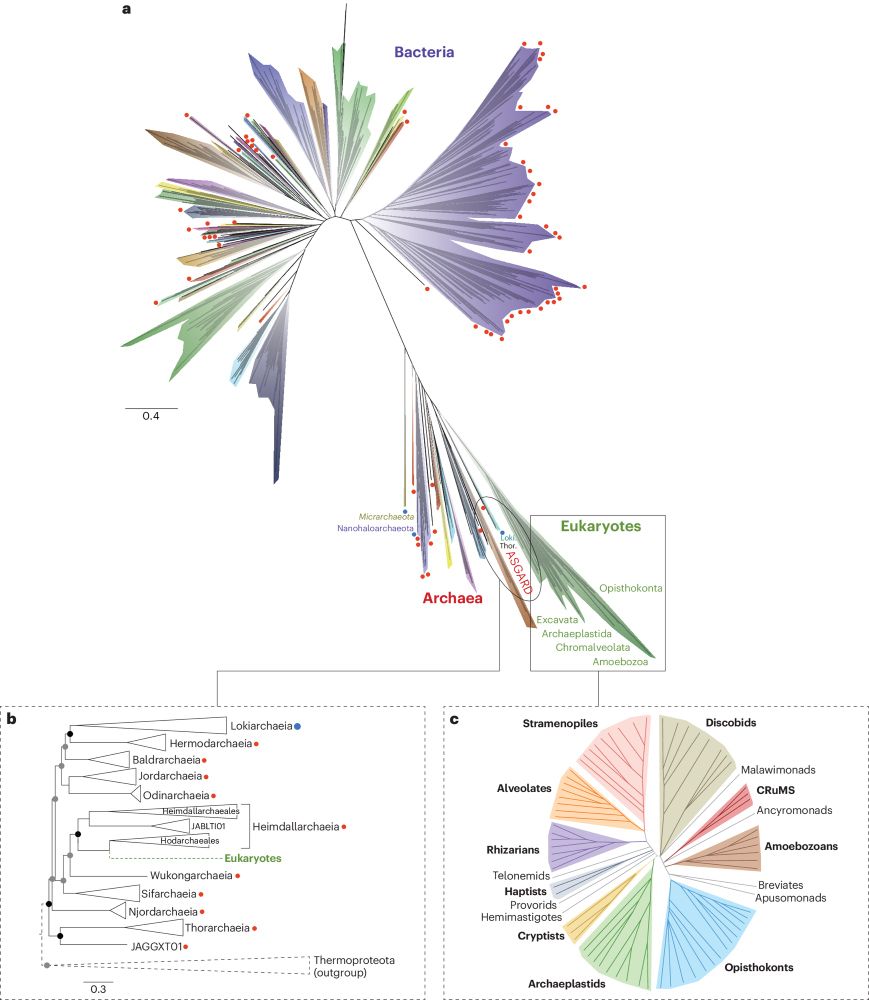
Nature Microbiology - The tree of life is a galvanizing image, anchoring biological diversity within a common framework. From a new view in 2016, the tree has continued to grow, and with it, our...
The so-called "VGIX01" bacterial phylum from GTDB now has a name :-) First paper from my collaboration with Lei Su, Jiangtao Li, Andreas Teske, and others. Lei spent a year in Aarhus getting started on this work as part of her PhD - more papers to come! journals.asm.org/doi/10.1128/...
The newly discovered bacterial phylum Candidatus Effluviviacota is widespread across diverse seepage ecosystems, marine environments, and freshwater environments, with a notable preference for cold se...