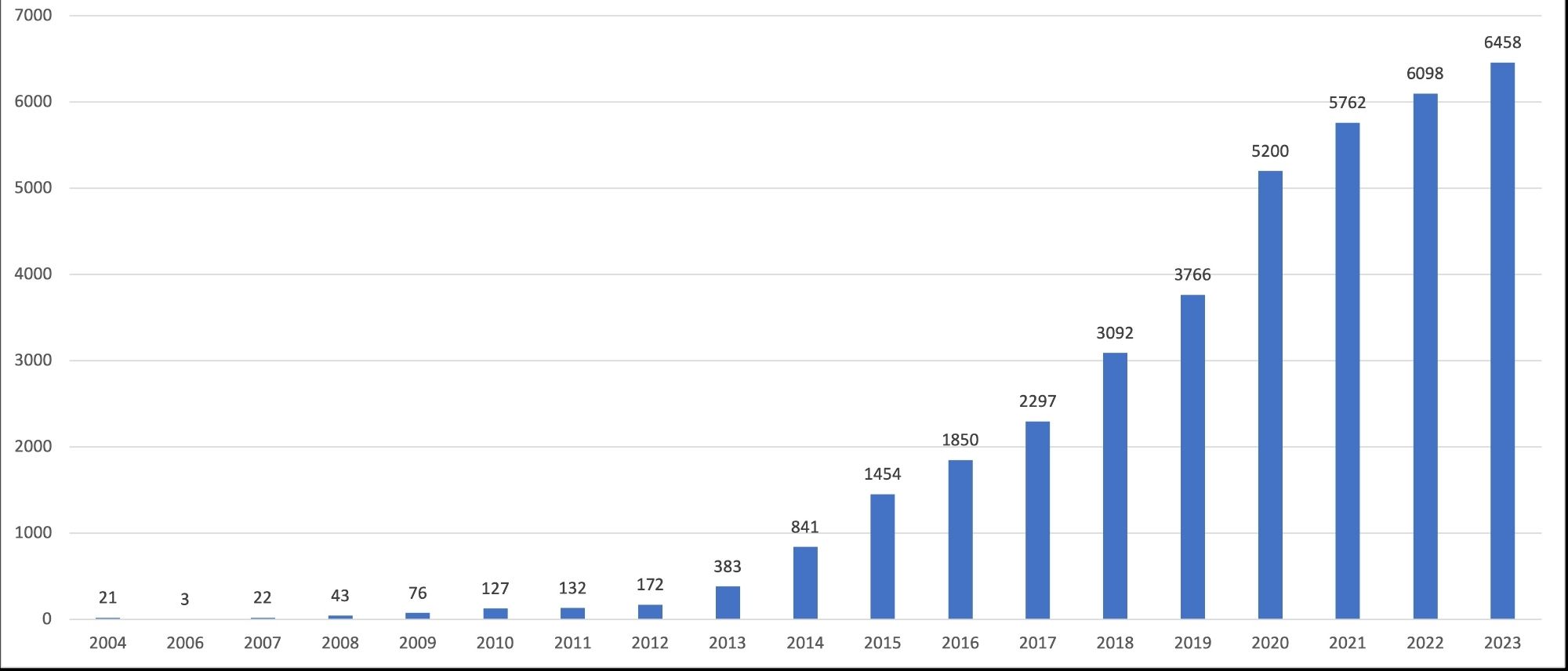
Since the release of the pride chatbot has been released we have seen more than 40 questions well responded to, no need for curator intervention. Now we don't have to reply to questions like: How can I download the entire PRIDE?
Training in bad data only produces bad models. I have seen a lot of deep-learning models in proteomics and almost no research/talks about the data that was used for training.
I was not able to access it. Can you share a PDF link 🤔
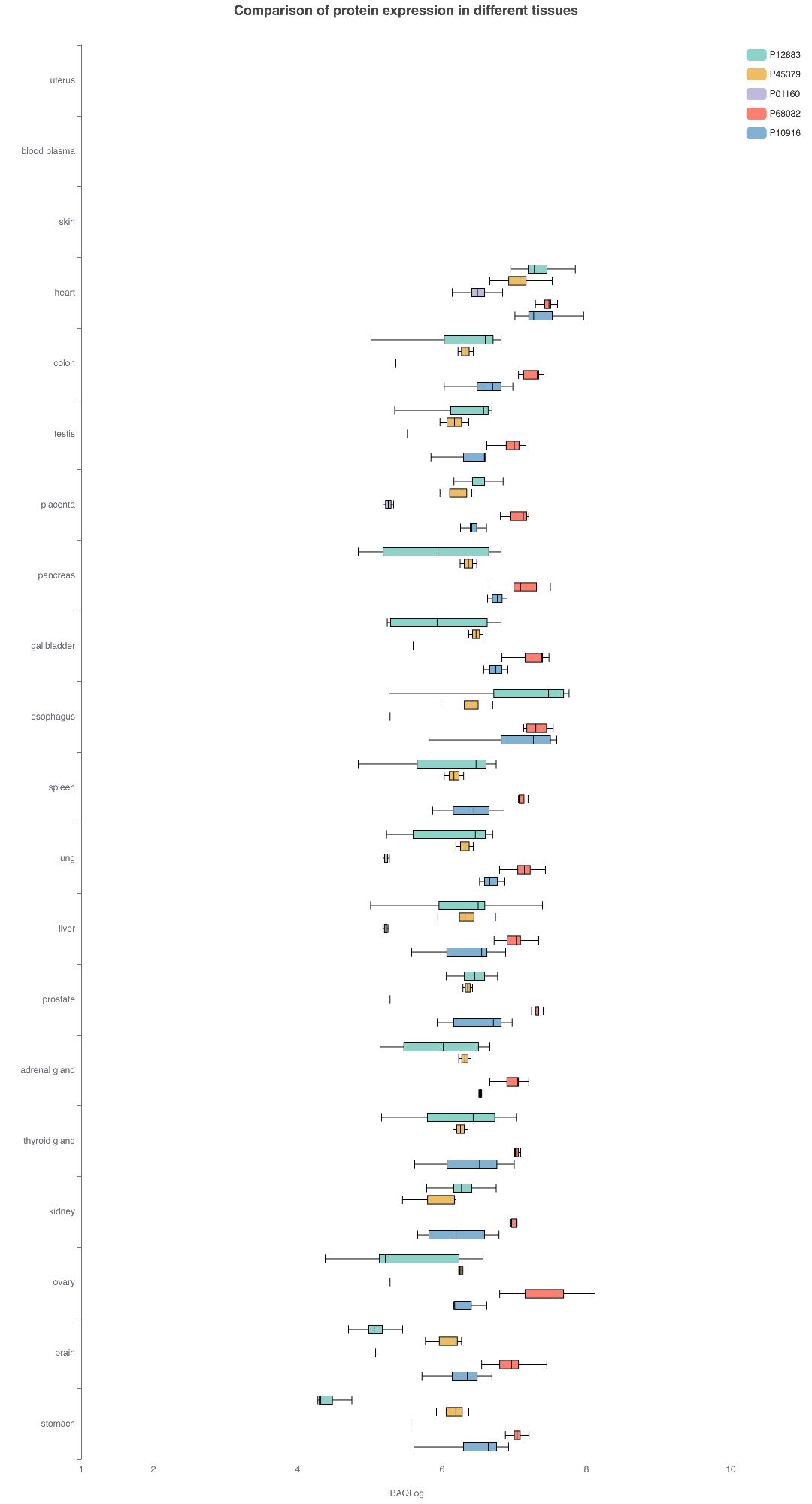
It is impressive how from all the proteins out there that have an expression profile that is what Uniprot called unambiguous "non-specific", it selected the 5 that are clearly "heart specific".
No, no, I think my preprint is given good quants for already well-known proteins. I'm also surprised how much chatgpt knows about simple questions. It is a really powerful tool for exploration, would love to training it with my data.
If you ask chatgpt to give you 5 proteins highly expressed in the human heart it responds: P12883, P45379, P01160, P68032, P10916. #quantmsquantms.org/ae/tissues?p...
Creazy idea: for those papers reanalysing large collections of datasets to put in the author list something like "Data Contributors", making easy for the authors that generate the original data to add the paper to their CV. Trying to give valuable Credit to the data submitters. What do you think?
We have developed a new python command-line tool to generate the PRIDE checksum files www.ebi.ac.uk/pride/markdo... this tool could help you to produce outside the PX submission tool your checksum.txt, nice for big submissions. Our target in the future is those submissions
Nice introduction in Chinese about quantms and what you can do with it in the quantms channel. www.youtube.com/watch?v=w3_b...#proteomics#massspectrometry#bigdata