
The RAT is out! 🐀 Our new Read Annotation Tool integrates taxonomic signals from MAGs, contigs, and reads from #metagenomicswww.nature.com/articles/s41.... 🧵 (1/5)
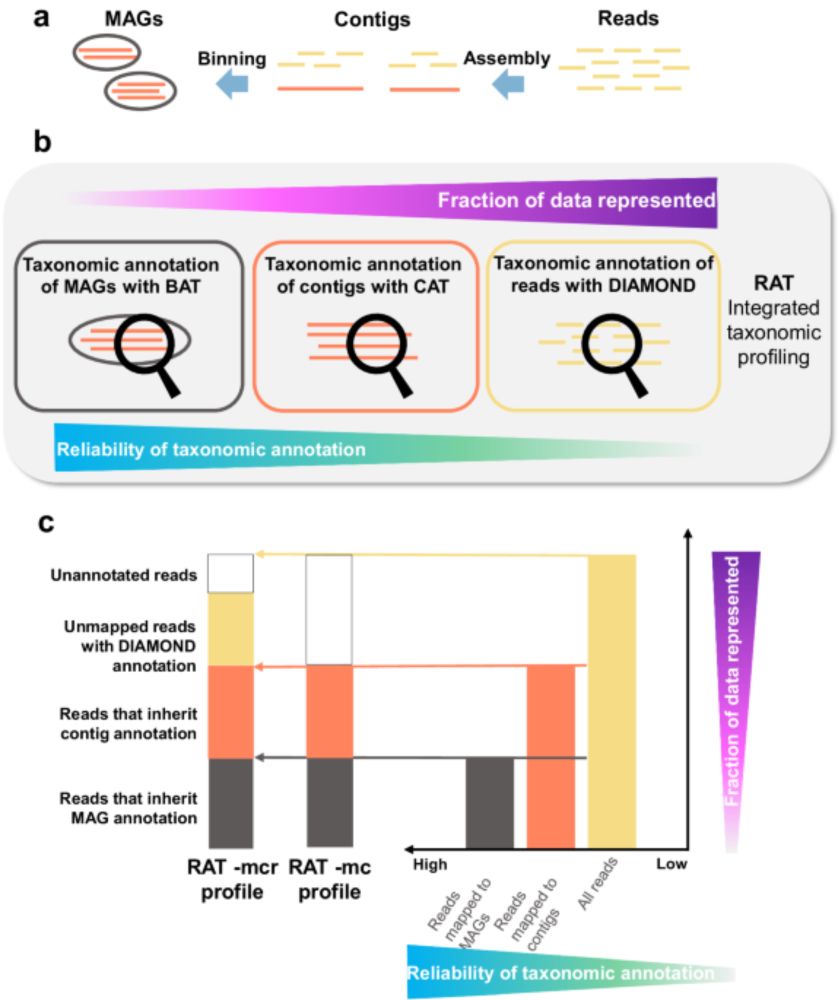
Metagenomic taxonomic profiling usually relies either on reads or assembled contigs/MAGs. Here, authors present RAT, a tool that integrates taxonomic signals from reads, contigs, and MAGs into one pro...
Do you want to know which virus discovery tool to use? Check out @lingyi.bsky.social 's new paper:
#Viromics #VirusDiscovery 🦠 by #Metagenomics 🧬! We used real metagenomic data from diverse biomes to #benchmark state-of-the-art #bioinformatics virus identification tools: genomebiology.biomedcentral.com/articles/10.... @goncalopiedade.bsky.social @bedutilh.bsky.social 🧵 (1/4)
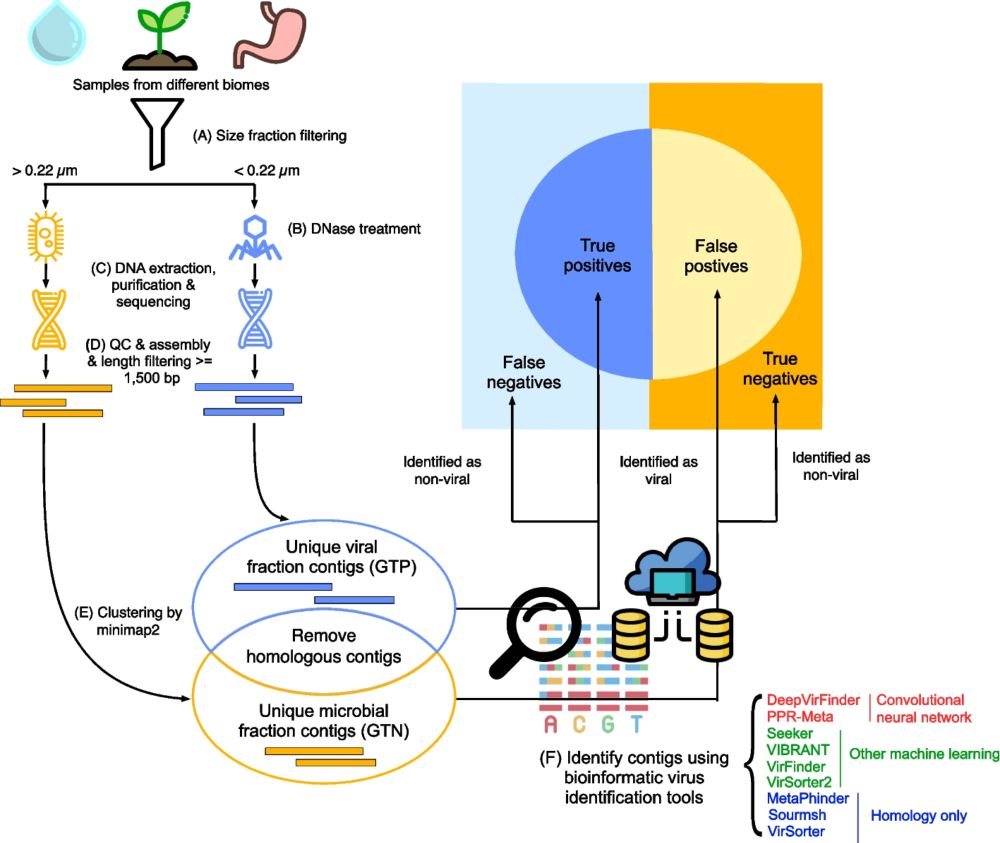
Background As most viruses remain uncultivated, metagenomics is currently the main method for virus discovery. Detecting viruses in metagenomic data is not trivial. In the past few years, many bioinfo...
#Viromics#VirusDiscovery#Metagenomics#benchmark#bioinformaticsgenomebiology.biomedcentral.com/articles/10....@goncalopiedade.bsky.social@bedutilh.bsky.social 🧵 (1/4)

Background As most viruses remain uncultivated, metagenomics is currently the main method for virus discovery. Detecting viruses in metagenomic data is not trivial. In the past few years, many bioinfo...
Excited to share the last chapter of #XinZhang'sbiorxiv.org/content/10.1...@binfutrecht.bsky.social#UUBeta#WesterdijkBiodiversity
Happy to see our most recent work on genome evolution of Fusarium isolates infecting banana published in @NewPhyt nph.onlinelibrary.wiley.com/doi/10.1111/...#AvWesterhoven@binfutrecht.bsky.social#UUBeta#WURplant#PhytoWUR@keygene.com
TBB Paper alert! Sam (@derdunk.bsky.social) shows how genomes evolve intracellular signaling when enforcing endosymbiosis, and how this resolves the conflict between the two players!
The last chapter of my PhD is out now! #PLOSCompBio: Intracellular signaling in proto-eukaryotes evolves to alleviate regulatory conflicts of endosymbiosis t.co/IUCEnNcBkG
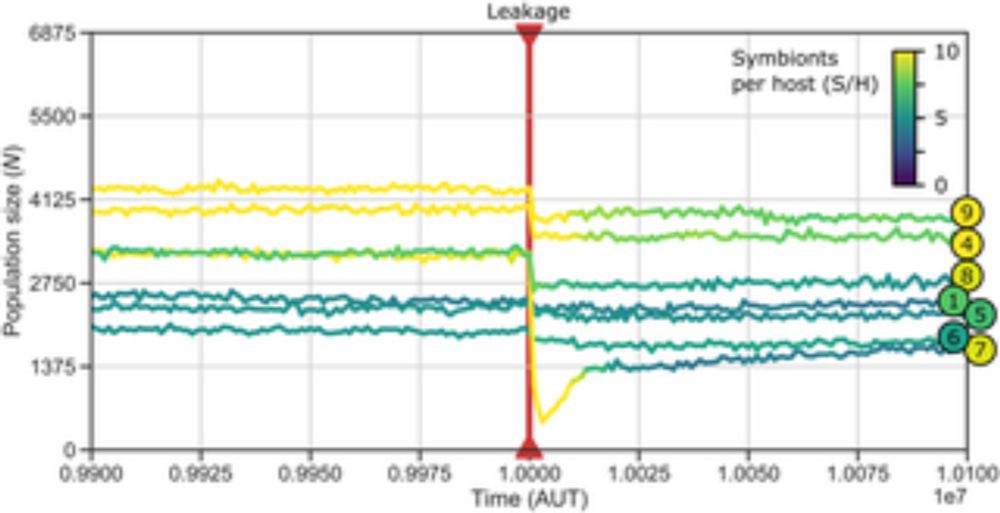
Author summary Virtually all life forms visible by the naked eye are eukaryotes, complex cells with a nucleus, mitochondria and many other subcellular structures as well as the regulatory mechanisms t...
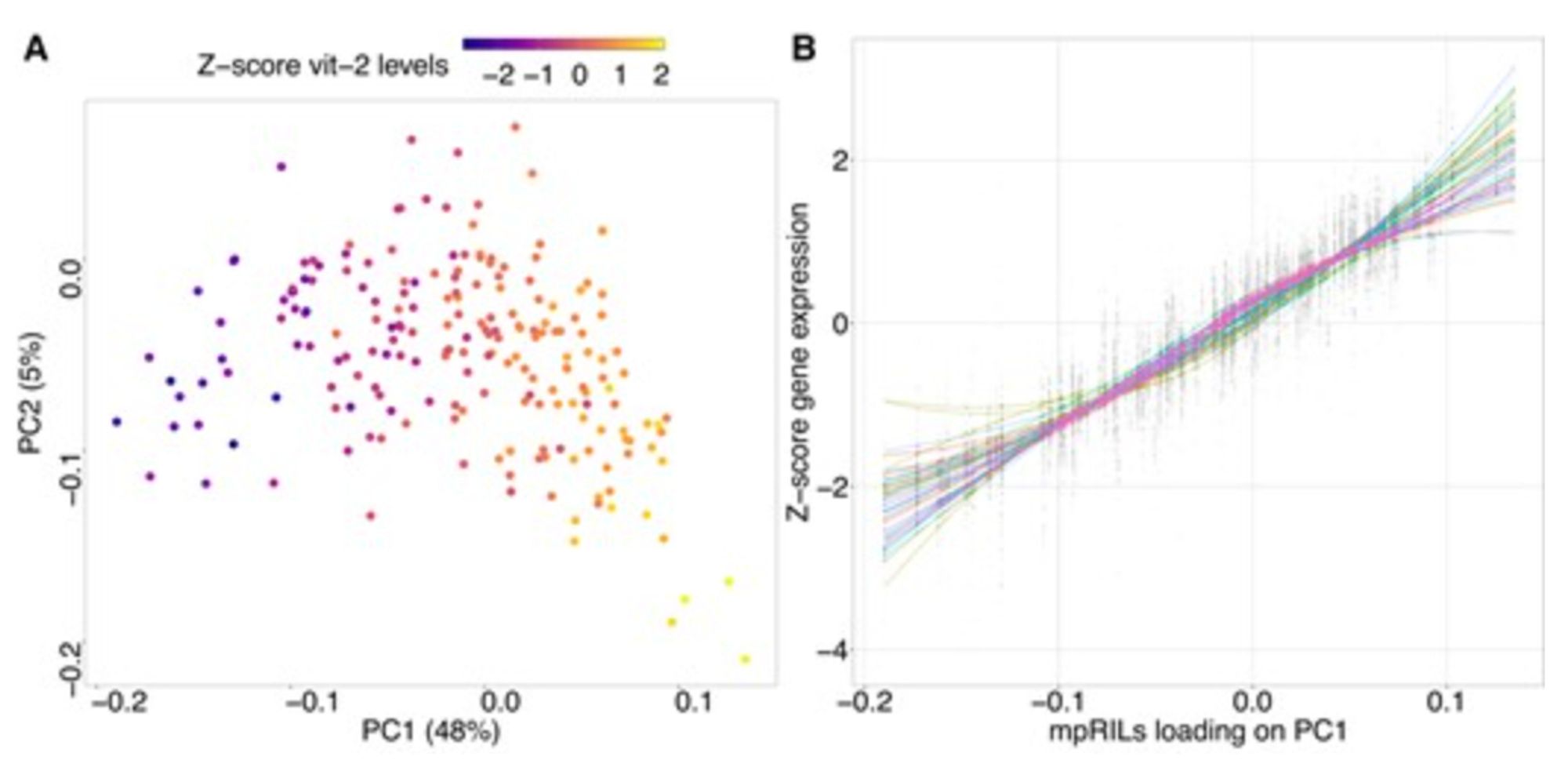
Paper alert: Van Eijnatten and colleagues find that C. elegans gene expression is confounded by what they call 'developmental age' (not chronological age). This complicates the interpretation of eQTL experiments, for which they provide a solution!
With a complex developmental program, eQTL mapping can be challenging, as changes in gene expression complicate interpretation. Check out this new paper in G3 by Bram van Eijnatten, Basten Snoek and collaborators from Wageningen University & Research, and see how we can improve detection of eQTLs!
Q&A with Paulien Hogeweg #NatureComputationalSciencewww.nature.com/articles/s43...

