Thanks for the replies. Social media win: Lauric Ferrat showed me the way: bsky.app/profile/lcpi...
I think the problem was I was trying with the new "fine-grained tokens" rather than the classic "personal access tokens" -- the below works fine for private repos! system("git clone [https://]<personal_access_token>@github.com/repo.git") Many thanks to Lauric Ferrat for pointing this out to me
Great idea. Honour to be included! Thank you. BSky really taking off now
It takes <30 seconds on my Linux server to go from a list of diagnostic codes to "date first diagnosed, any source" phenotype. It takes longer on Windows due to slower grepping of text files (~4 minutes). On the RAP it takes ~5 minutes to get the data from spark
Lots of caveats regarding the use of polygenic scores in non-EUR groups, but we aimed to apply the principles outlined in the recent report to the best of our current abilities, but future studies should move away from discrete ancestry clusters doi.org/10.17226/26902

Read online, download a free PDF, or order a copy in print or as an eBook.
Yes. Twitter had it's failings but I saw more relevant papers outside my direct followers then here, I presume due to "the algorithm" pairing me to similar posts/topics etc.
There seems to be a paper for every combination these days. I'm not saying all are bad - there are some good ones, and I review the ones in my area when I have the time - but many seem very superficial "We get these public summary stats for these two traits, got the IVW estimate, and here's a paper"
Time to invite some stats people!
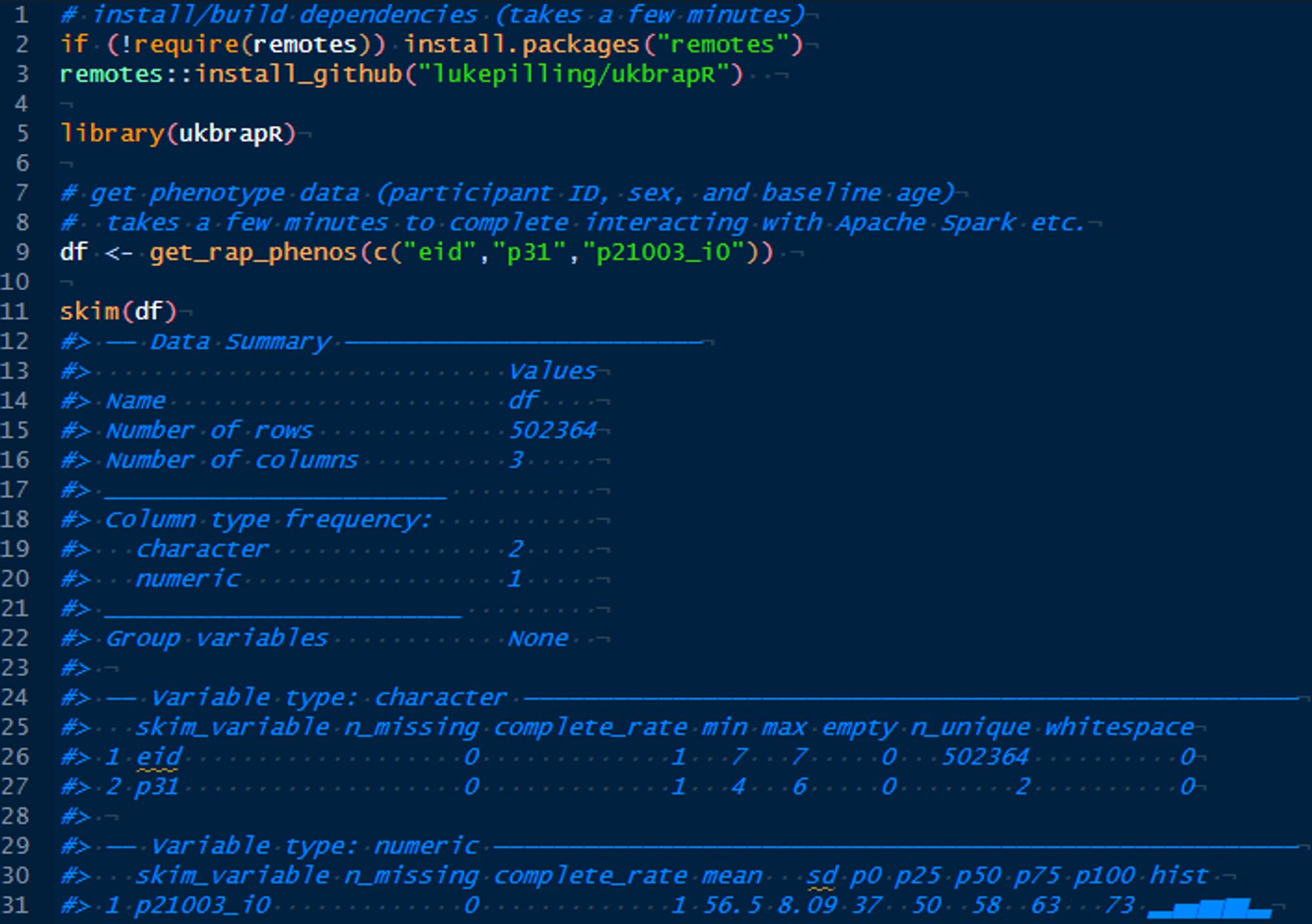
Once in a JupyterLab environment on a Spark Cluster the following R commands pull the data straight from the database without intermediate files Takes ~5 minutes to build all the package dependencies, and <5 minutes to get the data object