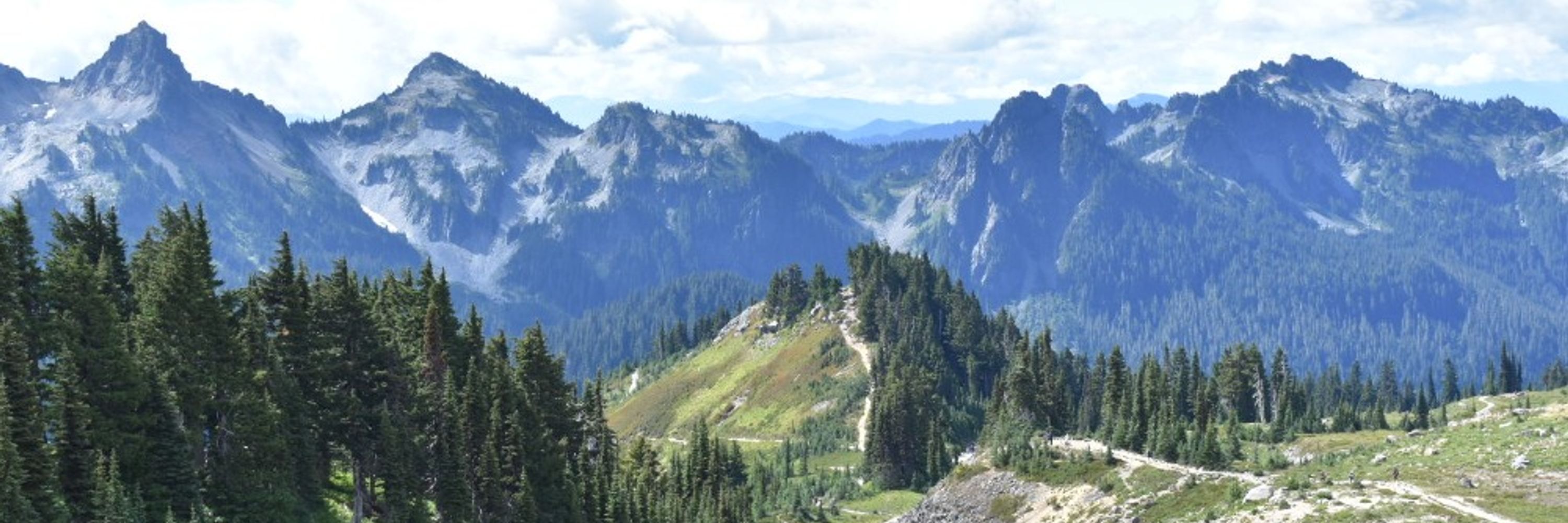

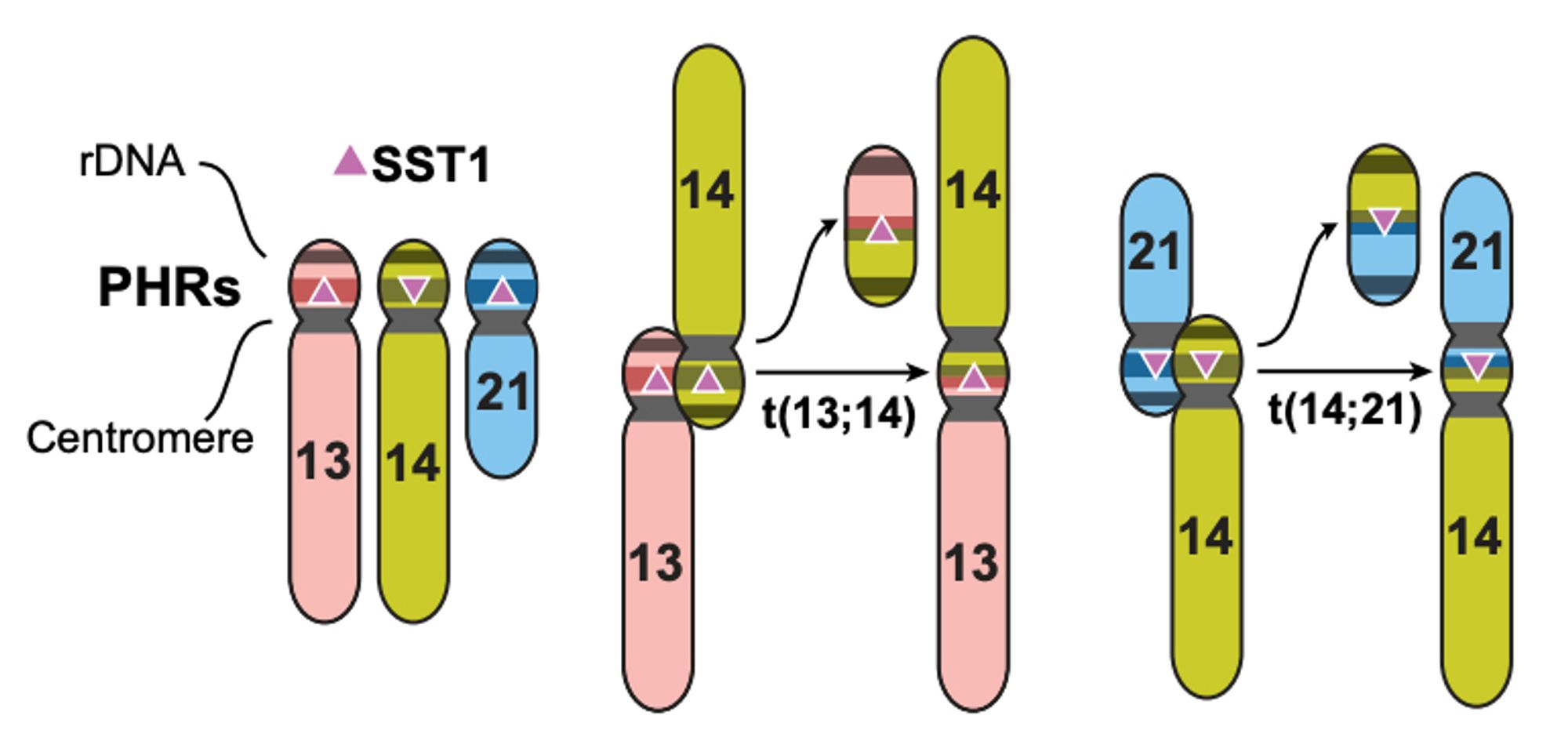
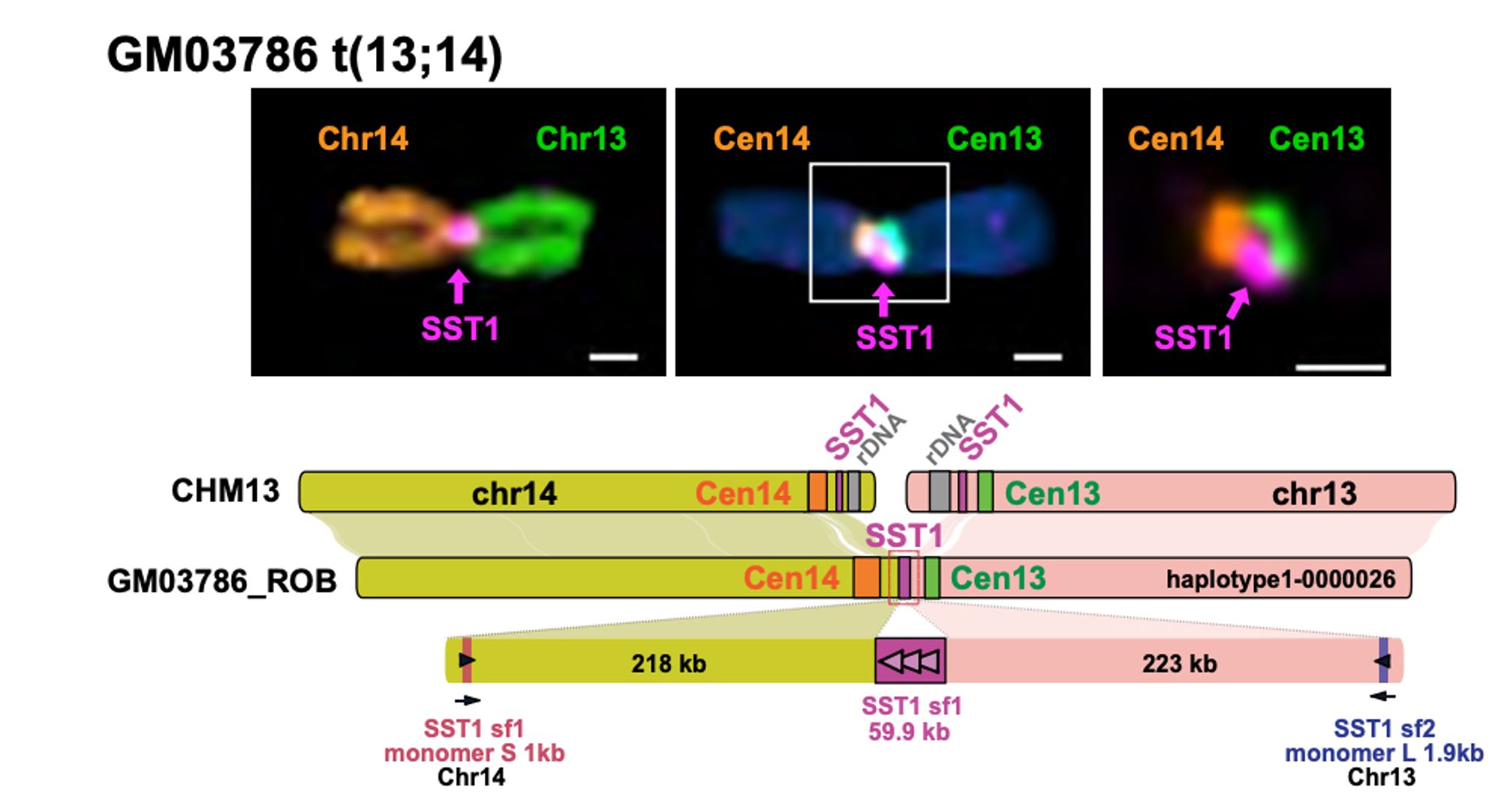
"The formation and propagation of human Robertsonian chromosomes" ROBs are the most common translocation in humans, approx. ~1 per 800 people. Last year we proposed a simple mechanism for ROB formation; This year we prove it by finishing some T2T! New preprint: www.biorxiv.org/content/10.1...
Calling all #virologypsu.wd1.myworkdayjobs.com/en-US/PSU_Ac...

APPLICATION INSTRUCTIONS: CURRENT PENN STATE EMPLOYEE (faculty, staff, technical service, or student), please login to Workday to complete the internal application process. Please do not apply here, a...
I'm hiring! Currently accepting applications for center coordinator, bioinformatics engineer/scientist, and postdoctoral researcher. Please share with those that might be interested: genomeinformatics.github.io/jobs2024/
Ruminant T2T paper is out! This is an open collaborative effort aiming to generate complete assemblies for agriculturally important and endangered ruminants and perform genomic analyses. Reach out to me if you are interested to join the immunogenomics working group! www.nature.com/articles/s41...
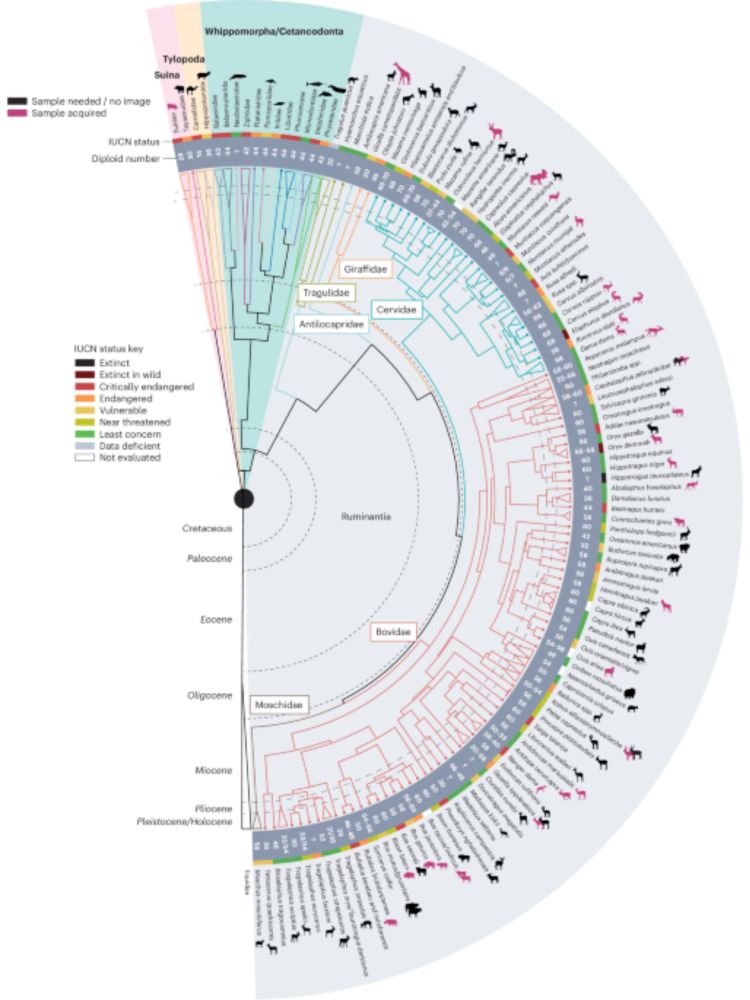
Here we describe an open collaborative effort termed the ‘Ruminant T2T Consortium’. It aims to generate complete diploid assemblies for many species of ruminants to examine chromosomal evolution in th...
Happy to be part of this effort! We compared IG and TR loci across four apes and revealed rapidly evolving regions containing species-specific immune genes. This suggests an incredible variation of adaptive immune responses across closely related species. Check out the preprint for more stories!
Check out our preprint describing a tool for reassembly of IG loci from LCLs containing somatic haplotypes produced by V(D)J recombination. We applied IgLoo to HPRC data and recovered many germline IG genes. We also revealed novel V(D)J recombination scenarios that might be confused with SVs
Check out our new preprint where we assessed the assembly quality of IG loci in nearly finished vertebrate genomes, classified systematic errors and provided examples on how to fix these errors using curated and transparent genome assembly
Unveiling Assembly Errors in Immunoglobulin Loci: A Comprehensive Evaluation of Long-read Genome Assemblies Across Vertebrates https://www.biorxiv.org/content/10.1101/2024.07.19.604360v1

Long-read sequencing technologies have revolutionized genome assembly, addressing the limitations of
Eric Engelbrecht in my lab showing us that we have much to discover in the human IGK locus! Gene conversions, novel inversions, SVs, SNVs, genes and alleles...oh my! rdcu.be/dJ8mL
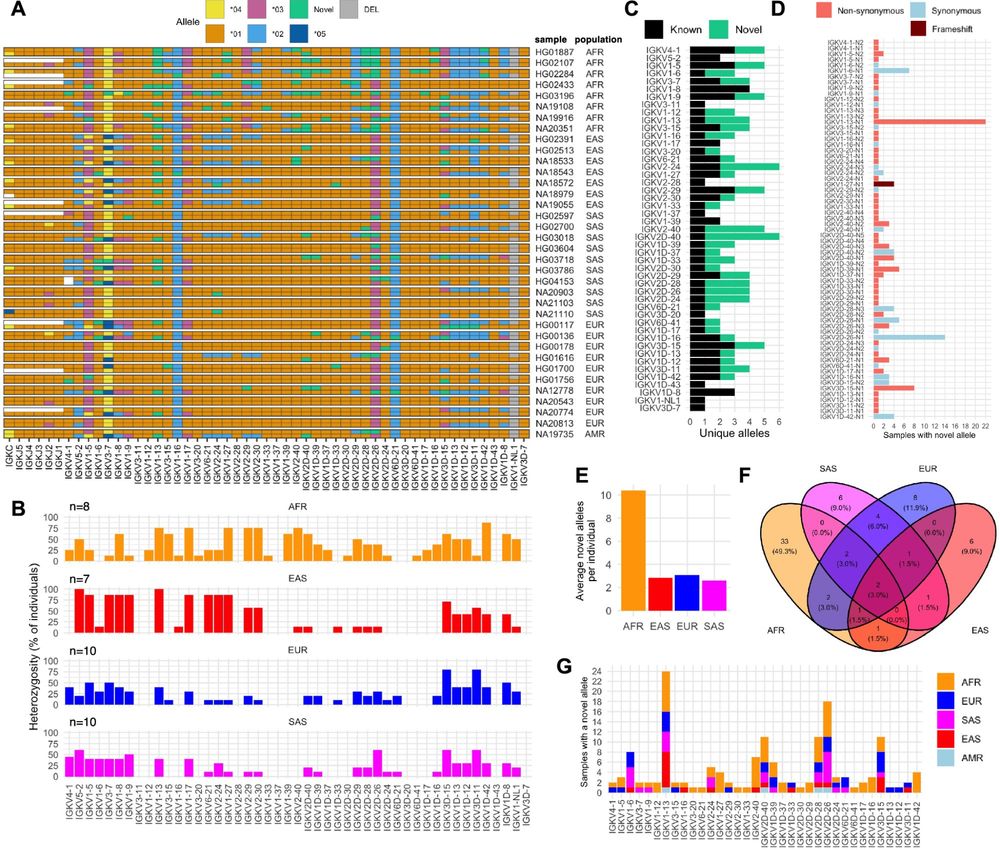
Genes & Immunity - Resolving haplotype variation and complex genetic architecture in the human immunoglobulin kappa chain locus in individuals of diverse ancestry
Skyview post here: skyview.social?url=https://...
Structural polymorphism and diversity of human segmental duplications www.biorxiv.org/content/10.1101/2024.06.04.597452v1

Segmental duplications (SDs) contribute significantly to human disease, evolution, and diversity yet
