
Funding: UKHSA, HPRU in HCAI & AMRin collaboration with the University of Oxford And many discussions this summer at Kavli Institute in Santa Barbara!
This was an amazing within UKHSA collaboration, with Fan and Leo, who are data scientists with a statistical background, doing an amazing job at learning and understanding bioinfomatics and plasmid epidemiology! Any feedback/questions really appreciated!
What we DID test is that the within patient plasmid genetic distance is significantly different from the between plasmid genetic distance. Pointing to plasmid found within each patient not being randomly extracted from the dataset (hence probably spread by conjugation)
We cannot say whether having identical plasmids in different hosts in the same patient is the result of conjugation within that patient or a wide transmission bottleneck with several bacterial hosts. Still, a lot of conjugation happened somewhere...
In fact 83 of the 115 patients we had data for had at least two identical plasmids in different bacterial hosts. And ONLY TWO patients had sequences from both clades, which is a clear hint of separate acquisition: PAT229 (RED) and PAT219 (GREEN)

We obtained a tree of the plasmids. It clearly has 2 main branches: a diverse one, mainly sampled from London, and an epidemic one, mainly sampled from the North West. Most of the samples in the epidemic clade are identical or very similar.
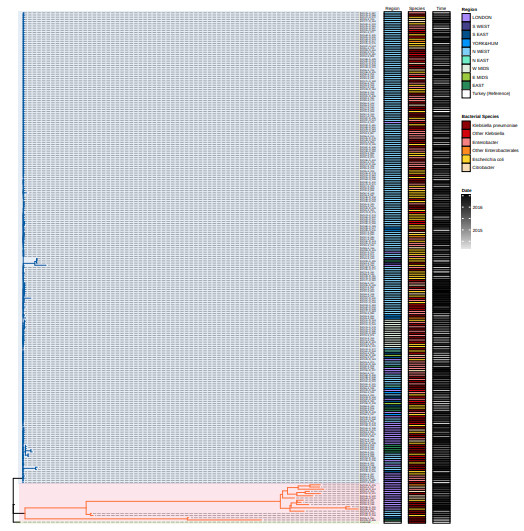
We used a sub-dataset from the from this paper, www.microbiologyresearch.org/content/jour... Selected sequences from patients who had at least 2 sequences, one of each carried an OXA48 resistance (Well known to be plasmid borne).

Introduction. Increasing numbers of carbapenemase-producing Enterobacterales (CPE), which can be challenging to treat, have been referred to the national reference laboratory in England since the ear...
First off: why do we care? In a plasmid outbreak, thanks to plasmid conjugation, we can find the same plasmid in different bacterial hosts and this could muddle our uncovering of the outbreak. Understanding how often this happens can help us devise appropriate screening.
Finally this draft is out! The question is: how often do we find the same plasmid in different bacterial hosts in the same patient? Spoiler alert: in this dataset very often!
Plasmid conjugation drives within-patient plasmid diversity https://www.biorxiv.org/content/10.1101/2024.09.27.615342v1

Plasmids are well known vehicles of antimicrobial resistance (AMR) genes dissemination. Through conj

